(Part III) Benchmarking scAtlasVAE with another methods
Contents
(Part III) Benchmarking scAtlasVAE with another methods¶
This notebook contains codes for benchmarking scAtlasVAE.
Please check the package version at https://github.com/WanluLiuLab/scAtlasVAE/blob/master/environment.yml for reproducing the results.
For more information about the scAtlasVAE model, please see https://scatlasvae.readthedocs.io/en/latest/.
For retrieving datasets, please see https://zenodo.org/records/10472914.
Initializing Benchmarking Environment¶
#########################################
# Initializing Benchmarking Environment #
#########################################
import scatlasvae
import scanpy as sc
import scvi # scvi==1.0.4
import scarches #
import scib
import scanorama
import celltypist
# import packages
import scanpy as sc # import scanpy
import matplotlib
import matplotlib.pyplot as plt # import matplotlib
import numpy as np # import numpy
import pandas as pd # import pandas
import gc # import garbage collector
from typing import Literal, Union # import typing
import time
# set plot linewidth
def setPltLinewidth(linewidth:float): # define function to set plot linewidth
matplotlib.rcParams['axes.linewidth'] = linewidth # set plot linewidth
setPltLinewidth(1) # set plot linewidth to 1
# set plot parameters
plt.rcParams['figure.dpi'] = 300 # set figure resolution
plt.rcParams['savefig.dpi'] = 300 # set figure resolution
plt.rcParams['font.size'] = 8 # set font size
plt.rcParams['axes.linewidth'] = 1 # set plot linewidth
plt.rcParams['font.family'] = 'Arial' # set font family
# Load dataset
adata_cd8 = sc.read_h5ad('huARdb_v2_GEX.CD8.hvg4k.h5ad')
adata_cd8_zheng = sc.read_h5ad('adata_cd8_zheng.h5ad')
adata_cd8_chu = sc.read_h5ad('adata_cd8_chu.h5ad')
adata_cd8_multi_atlas = sc.read_h5ad(
'huARdb_v2_GEX.CD8.hvg4k.pan_cancer_multi_atlas_vae.h5ad'
)
# Useful functions
try:
import seaborn as sns
import matplotlib
import matplotlib.pyplot as plt
matplotlib.rcParams['font.family'] = 'Arial'
matplotlib.rcParams['font.size'] = '10'
matplotlib.rcParams['font.weight'] = 100
matplotlib.rcParams['axes.linewidth'] = 2
matplotlib.rcParams['axes.edgecolor'] = "#000000"
def createFig(figsize=(8, 4)):
fig,ax=plt.subplots()
ax.spines['right'].set_color('none')
ax.spines['top'].set_color('none')
#ax.spines['bottom'].set_color('none')
#ax.spines['left'].set_color('none')
for line in ax.yaxis.get_ticklines():
line.set_markersize(5)
line.set_color("#585958")
line.set_markeredgewidth(0.5)
for line in ax.xaxis.get_ticklines():
line.set_markersize(5)
line.set_markeredgewidth(0.5)
line.set_color("#585958")
ax.set_xbound(0,10)
ax.set_ybound(0,10)
fig.set_size_inches(figsize)
return fig,ax
def createSubplots(nrow,ncol, figsize=(8,8),gridspec_kw={}):
fig,axes=plt.subplots(nrow, ncol, gridspec_kw=gridspec_kw)
for ax in axes.flatten():
ax.spines['right'].set_color('none')
ax.spines['top'].set_color('none')
for line in ax.yaxis.get_ticklines():
line.set_markersize(5)
line.set_color("#585958")
line.set_markeredgewidth(0.5)
for line in ax.xaxis.get_ticklines():
line.set_markersize(5)
line.set_markeredgewidth(0.5)
line.set_color("#585958")
fig.set_size_inches(figsize)
return fig,axes
hist_kws={"linewidth": 0, "alpha": .5}
except:
print("failed to load plotting packages")
Main Benchmarking Script¶
Benchmark Zheng et al., 2019 Dataset¶
Benchmarking integration performance¶
##############
# scAtlasVAE #
##############
vae_model = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_zheng,
batch_key=["sample_name", "study_name"],
batch_embedding="embedding",
batch_hidden_dim=64,
device="cuda:0",
)
vae_model.fit(max_epoch=10, lr=5e-5)
adata_cd8_zheng.obsm["X_gex"] = vae_model.get_latent_embedding(show_progress=True)
vae_model = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_zheng,
batch_key=["sample_name", "study_name"],
batch_embedding="embedding",
batch_hidden_dim=64,
label_key="cell_subtype_zheng_2021",
device="cuda:0",
)
vae_model.fit(max_epoch=10, lr=5e-5)
adata_cd8_zheng.obsm["X_gex_supervised"] = vae_model.get_latent_embedding(
show_progress=True
)
###################
# scVI and scANVI #
###################
scvi.model.SCVI.setup_anndata(
adata_cd8_zheng, batch_key="sample_name", categorical_covariate_keys=["study_name"]
)
model = scvi.model.SCVI(adata_cd8_zheng)
model.train(max_epochs=10)
adata_cd8_zheng.obsm["X_scVI"] = model.get_latent_representation()
scvi.model.SCANVI.setup_anndata(
adata_cd8_zheng,
batch_key="sample_name",
categorical_covariate_keys=["study_name"],
labels_key="cell_subtype_zheng_2021",
unlabeled_category="undefined",
)
model = scvi.model.SCANVI(adata_cd8_zheng)
model.train(max_epochs=10)
adata_cd8_zheng.obsm["X_scANVI"] = model.get_latent_representation()
##########
# scPoli #
##########
scpoli_model = scarches.models.scpoli.scPoli(
adata=adata_cd8_zheng,
condition_keys=["sample_name", "study_name"],
recon_loss="zinb",
)
early_stopping_kwargs = {
"early_stopping_metric": "val_prototype_loss",
"mode": "min",
"threshold": 0,
"patience": 20,
"reduce_lr": True,
"lr_patience": 13,
"lr_factor": 0.1,
}
scpoli_model.train(
n_epochs=10,
pretraining_epochs=5,
early_stopping_kwargs=early_stopping_kwargs,
eta=5,
)
for i in tqdm.trange(0, len(adata_cd8_zheng), 320):
Z.append(scpoli_model.get_latent(adata_cd8_zheng[i : i + 320]))
adata_cd8_zheng.obsm["X_scPoli"] = Z
scpoli_model = scarches.models.scpoli.scPoli(
adata=adata_cd8_zheng,
condition_keys=["sample_name", "study_name"],
recon_loss="zinb",
cell_type_keys=["cell_subtype_zheng_2021"],
unknown_ct_names=["undefined"],
)
scpoli_model.train(
n_epochs=10,
pretraining_epochs=5,
early_stopping_kwargs=early_stopping_kwargs,
eta=5,
)
Z = []
import tqdm
# scpoli_model.
for i in tqdm.trange(0, len(adata_cd8_zheng), 320):
Z.append(scpoli_model.get_latent(adata_cd8_zheng[i : i + 320]))
adata_cd8_zheng.obsm["X_scPoli_supervised"] = Z
##########
# SCALEX #
##########
os.chdir("~/Biosoft/SCALEX")
from scalex import SCALEX
adata_cd8_zheng.obsm["X"] = adata_cd8_zheng.X
adata_cd8_zheng_scalex = SCALEX(
adata_cd8_zheng,
batch_name=["study_name"],
n_top_features=adata_cd8_zheng.shape[1],
min_cells=1,
min_features=1,
use_layer="X",
ignore_umap=True,
)
adata_cd8_zheng.obsm["X_SCALEX"] = adata_cd8_zheng_scalex.obsm["latent"]
###########
# Harmony #
###########
from harmony import harmonize
sc.pp.normalize_total(adata_cd8_zheng)
sc.pp.log1p(adata_cd8_zheng)
sc.tl.pca(adata_cd8_zheng)
adata_cd8_zheng.obsm["X_harmony"] = harmonize(
adata_cd8_zheng.obsm["X_pca"], adata_cd8_zheng.obs, batch_key="study_name"
)
#############
# Scanoarma #
#############
def scanoramaCorrectMerge(adata, groupby="sample_name"):
adatas = []
group_names = list(np.unique(adata.obs[groupby]))
for i in group_names:
adatas.append(adata[adata.obs[groupby] == i])
corrected = scanorama.correct_scanpy(adatas, dimred=10, return_dimred=True)
return sc.concat(corrected)[adata.obs.index]
adata_cd8_zheng.obsm["X_scanorama"] = scanoramaCorrectMerge(
adata_cd8_zheng, groupby="study_name"
).obsm["X_scanorama"]
silhouette_batch_result = {"sample_name": {}, "study_name": {}}
for emb_key in [
"X_gex",
"X_gex_supervised",
"X_scANVI",
"X_scVI",
"X_scPoli",
"X_scPoli_supervised",
"X_pca",
"X_harmony",
"X_scanorama",
"X_DESC",
"X_SCALEX",
"X_RPCA",
"X_CCA",
]:
if emb_key in adata_cd8_zheng.obsm.keys():
adata_ = adata_cd8_zheng[
adata_cd8_zheng.obs["cell_subtype_zheng_2021"] != "undefined"
]
sc.pp.neighbors(adata_, use_rep=emb_key)
silhouette_batch_result["sample_name"][emb_key] = scib.me.silhouette_batch(
adata_,
batch_key="sample_name",
label_key="cell_subtype_zheng_2021",
embed=emb_key,
)
silhouette_batch_result["study_name"][emb_key] = scib.me.silhouette_batch(
adata_,
batch_key="study_name",
label_key="cell_subtype_zheng_2021",
embed=emb_key,
)
silhouette_result = {"study_name": {}}
for emb_key in [
"X_gex",
"X_gex_supervised",
"X_scANVI",
"X_scVI",
"X_scPoli",
"X_scPoli_supervised",
"X_pca",
"X_harmony",
"X_scanorama",
"X_DESC",
"X_SCALEX",
]:
if emb_key in adata_cd8_zheng.obsm.keys():
adata_ = adata_cd8_zheng[
adata_cd8_zheng.obs["cell_subtype_zheng_2021"] != "undefined"
]
sc.pp.neighbors(adata_, use_rep=emb_key)
silhouette_result["study_name"][emb_key] = scib.me.silhouette(
adata_, label_key="cell_subtype_zheng_2021", embed=emb_key
)
graph_connectivity_result = {}
for emb_key in [
"X_gex",
"X_gex_supervised",
"X_scANVI",
"X_scVI",
"X_scPoli",
"X_scPoli_supervised",
"X_pca",
"X_harmony",
"X_scanorama",
"X_DESC",
"X_SCALEX",
"X_RPCA",
"X_CCA",
]:
if emb_key in adata_cd8_zheng.obsm.keys():
adata_ = adata_cd8_zheng[
adata_cd8_zheng.obs["cell_subtype_zheng_2021"] != "undefined"
]
sc.pp.neighbors(adata_, use_rep=emb_key)
graph_connectivity_result[emb_key] = scib.me.graph_connectivity(
adata_,
label_key="cell_subtype_zheng_2021",
)
pcr_comparison_result = {"sample_name": {}, "study_name": {}}
for emb_key in [
"X_gex",
"X_gex_supervised",
"X_scANVI",
"X_scVI",
"X_scPoli",
"X_scPoli_supervised",
"X_pca",
"X_harmony",
"X_scanorama",
"X_RPCA",
"X_CCA",
"X_SCALEX",
]:
if emb_key in adata_cd8_zheng.obsm.keys():
adata_ = adata_cd8_zheng[
adata_cd8_zheng.obs["cell_subtype_zheng_2021"] != "undefined"
]
# sc.pp.neighbors(adata_, use_rep=emb_key)
pcr_comparison_result["study_name"][emb_key] = scib.me.pcr_comparison(
adata_, adata_, covariate="study_name", embed=emb_key, n_comps=10
)
pcr_comparison_result["sample_name"][emb_key] = scib.me.pcr_comparison(
adata_, adata_, covariate="sample_name", embed=emb_key, n_comps=10
)
isolated_labels_asw_result = {"sample_name": {}, "study_name": {}}
for emb_key in [
"X_gex",
"X_gex_supervised",
"X_scANVI",
"X_scVI",
"X_scPoli",
"X_scPoli_supervised",
"X_pca",
"X_harmony",
"X_scanorama",
"X_DESC",
"X_SCALEX",
"X_RPCA",
"X_CCA",
]:
if emb_key in adata_cd8_zheng.obsm.keys():
adata_ = adata_cd8_zheng[
adata_cd8_zheng.obs["cell_subtype_zheng_2021"] != "undefined"
]
sc.pp.neighbors(adata_, use_rep=emb_key)
isolated_labels_asw_result["sample_name"][
emb_key
] = scib.me.isolated_labels_asw(
adata_,
label_key="cell_subtype_zheng_2021",
embed=emb_key,
batch_key="sample_name",
)
isolated_labels_asw_result["study_name"][emb_key] = scib.me.isolated_labels_asw(
adata_,
label_key="cell_subtype_zheng_2021",
embed=emb_key,
batch_key="study_name",
)
isolated_labels_f1_result = {"sample_name": {}, "study_name": {}}
for emb_key in [
"X_gex",
"X_gex_supervised",
"X_scANVI",
"X_scVI",
"X_scPoli",
"X_scPoli_supervised",
"X_pca",
"X_harmony",
"X_scanorama",
"X_DESC",
"X_SCALEX",
"X_RPCA",
"X_CCA",
]:
if emb_key in adata_cd8_zheng.obsm.keys():
adata_ = adata_cd8_zheng[
adata_cd8_zheng.obs["cell_subtype_zheng_2021"] != "undefined"
]
sc.pp.neighbors(adata_, use_rep=emb_key)
isolated_labels_f1_result["sample_name"][emb_key] = scib.me.isolated_labels_f1(
adata_,
label_key="cell_subtype_zheng_2021",
embed=emb_key,
batch_key="sample_name",
)
isolated_labels_f1_result["study_name"][emb_key] = scib.me.isolated_labels_f1(
adata_,
label_key="cell_subtype_zheng_2021",
embed=emb_key,
batch_key="study_name",
)
Benchmarking label transfer performance¶
Left 5% random cells from study¶
import sklearn
from sklearn.metrics import roc_auc_score
roc_auc_scores_scatlasvae_zero_shot = {
i: [] for i in [1.0, 2.0, 3.0, 4.0, 5.0, 6.0, 7.0]
}
for i in [1.0, 2.0, 3.0, 4.0, 5.0, 6.0, 7.0]:
for _ in range(3):
l = list(range(adata_cd8_zheng.shape[0]))
train, test = sklearn.model_selection.train_test_split(l, test_size=0.05)
adata_cd8_zheng_train = adata_cd8_zheng[train]
adata_cd8_zheng_test = adata_cd8_zheng[test]
vae_model = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_zheng_train,
batch_key=["sample_name", "study_name"],
batch_embedding="embedding",
batch_hidden_dim=64,
label_key="cell_subtype_zheng_2021",
device="cuda:0",
)
vae_model.fit(
max_epoch=10,
lr=5e-5,
pred_weight=i,
)
checkpoint_path = "./adata_cd8_zheng_models/left.model"
vae_model.save_to_disk(checkpoint_path)
import torch
state_dict = torch.load(checkpoint_path, map_location="cpu")
config = state_dict["model_config"]
vae_model_test = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_zheng_test,
pretrained_state_dict=state_dict["model_state_dict"],
**config,
)
from sklearn.metrics import roc_auc_score
vae_model_test.eval()
label_predictions = vae_model_test.predict_labels().detach().cpu().numpy()
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(list(map(lambda x: x[-2], vae_model_test._dataset)))[
[vae_model_test._shuffle_indices]
].unsqueeze(1),
vae_model_test.n_label + 1,
)
roc_auc_scores_scatlasvae_zero_shot[i].append(
roc_auc_score(y, label_predictions, average=None).mean()
)
roc_auc_scores_scatlasvae_full_shot = {
i: [] for i in [1.0, 2.0, 3.0, 4.0, 5.0, 6.0, 7.0]
}
for i in [1.0, 2.0, 3.0, 4.0, 5.0, 6.0, 7.0]:
for _ in range(10):
l = list(range(adata_cd8_zheng.shape[0]))
train, test = sklearn.model_selection.train_test_split(l, test_size=0.05)
adata_cd8_zheng.obs["cell_subtype_zheng_2021_scAtlasVAE"] = list(
adata_cd8_zheng.obs["cell_subtype_zheng_2021"]
)
adata_cd8_zheng.obs.iloc[
test,
list(adata_cd8_zheng.obs.columns).index(
"cell_subtype_zheng_2021_scAtlasVAE"
),
] = "undefined"
vae_model = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_zheng,
batch_key=["sample_name", "study_name"],
batch_embedding="embedding",
batch_hidden_dim=64,
label_key="cell_subtype_zheng_2021_scANVI",
device="cuda:0",
)
vae_model.fit(max_epoch=10, lr=5e-5, pred_weight=i)
from sklearn.metrics import roc_auc_score
vae_model.eval()
label_predictions = vae_model.predict_labels().detach().cpu().numpy()
col = list(vae_model.label_category.categories)
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(
list(
map(
lambda x: col.index(x),
adata_cd8_zheng.obs["cell_subtype_zheng_2021"],
)
)
).unsqueeze(1),
vae_model.n_label + 1,
)
test_set = set(test)
indices = np.array(
adata_cd8_zheng.obs["cell_subtype_zheng_2021"] != "undefined"
) & np.array(list(map(lambda x: x in test_set, range(len(adata_cd8_zheng)))))
sel = y[indices][:, :-1].sum(0) > 0
roc_auc_scores_scatlasvae_full_shot[i].append(
roc_auc_score(
y[indices][:, :-1][:, sel],
label_predictions[indices][:, sel],
average=None,
).mean()
)
scANVI_roc_auc_scores = []
for _ in range(10):
l = list(range(adata_cd8_zheng.shape[0]))
train, test = sklearn.model_selection.train_test_split(l, test_size=0.05)
adata_cd8_zheng.obs["cell_subtype_zheng_2021_scANVI"] = list(
adata_cd8_zheng.obs["cell_subtype_zheng_2021"]
)
adata_cd8_zheng.obs.iloc[
test, list(adata_cd8_zheng.obs.columns).index("cell_subtype_zheng_2021_scANVI")
] = "undefined"
scvi.model.SCANVI.setup_anndata(
adata_cd8_zheng,
batch_key="sample_name",
categorical_covariate_keys=["study_name"],
labels_key="cell_subtype_zheng_2021_scANVI",
unlabeled_category="undefined",
)
model = scvi.model.SCANVI(adata_cd8_zheng)
model.train(max_epochs=10)
scanvi_prediction = model.predict(soft=True)
col = list(scanvi_prediction.columns) + ["undefined"]
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(
list(
map(
lambda x: col.index(x),
adata_cd8_zheng.obs["cell_subtype_zheng_2021"],
)
)
).unsqueeze(1),
len(col),
)
test = set(test)
indices = list(
map(
lambda x: x[1] != "undefined" and x[0] in test,
enumerate(adata_cd8_zheng.obs["cell_subtype_zheng_2021"]),
)
)
scANVI_roc_auc_scores.append(
roc_auc_score(y[indices], scanvi_prediction.to_numpy()[indices])
)
scpoli_roc_auc_scores = []
for _ in range(10):
l = list(range(adata_cd8_zheng.shape[0]))
train, test = sklearn.model_selection.train_test_split(l, test_size=0.05)
adata_cd8_zheng_train = adata_cd8_zheng[train]
adata_cd8_zheng_test = adata_cd8_zheng[test]
scpoli_model = scarches.models.scpoli.scPoli(
adata=adata_cd8_zheng_train[
adata_cd8_zheng_train.obs["cell_subtype_zheng_2021"] != "undefined"
],
condition_keys=["sample_name", "study_name"],
cell_type_keys=["cell_subtype_zheng_2021"],
unknown_ct_names=["undefined"],
recon_loss="zinb",
)
early_stopping_kwargs = {
"early_stopping_metric": "val_prototype_loss",
"mode": "min",
"threshold": 0,
"patience": 20,
"reduce_lr": True,
"lr_patience": 13,
"lr_factor": 0.1,
}
scpoli_model.train(
n_epochs=10,
pretraining_epochs=5,
early_stopping_kwargs=early_stopping_kwargs,
eta=5,
prototype_training=True,
)
# adata_cd8_zheng_test.obs.pop('cell_subtype_zheng_2021')
# adata_cd8_zheng_test.obs['cell_subtype_zheng_2021'] = 'undefined'
scpoli_query = scarches.models.scpoli.scPoli.load_query_data(
adata=adata_cd8_zheng_test,
reference_model=scpoli_model,
labeled_indices=[],
)
scpoli_query.train(n_epochs=10, pretraining_epochs=5, eta=10)
c = list(scpoli_query.cell_types_.keys())
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(
list(
map(
lambda x: c.index(x) if x in c else vae_model_test.n_label,
adata_cd8_zheng_test.obs["cell_subtype_zheng_2021"],
)
)
).unsqueeze(1),
len(c),
)
results_dict = scpoli_query.classify(adata_cd8_zheng_test, scale_uncertainties=True)
indices = adata_cd8_zheng_test.obs["cell_subtype_zheng_2021"] != "undefined"
scpoli_roc_auc_scores.append(
roc_auc_score(
y[indices],
1 - results_dict["cell_subtype_zheng_2021"]["weighted_distances"][indices],
)
)
roc_auc_scores_cell_typist = []
for _ in range(10):
l = list(range(adata_cd8_zheng.shape[0]))
train, test = sklearn.model_selection.train_test_split(l, test_size=0.05)
adata_cd8_zheng_train = adata_cd8_zheng[train]
adata_cd8_zheng_train = adata_cd8_zheng_train[
adata_cd8_zheng_train.obs["cell_subtype_zheng_2021"] != "undefined"
]
adata_cd8_zheng_test = adata_cd8_zheng[test]
adata_cd8_zheng_test = adata_cd8_zheng_test[
adata_cd8_zheng_test.obs["cell_subtype_zheng_2021"] != "undefined"
]
sc.pp.normalize_total(adata_cd8_zheng_train, target_sum=1e4)
sc.pp.log1p(adata_cd8_zheng_train)
t_start = time.time()
model_fs = celltypist.train(
adata_cd8_zheng_train,
"cell_subtype_zheng_2021",
n_jobs=96,
max_iter=5,
use_SGD=True,
)
t_end = time.time()
print(f"Time elapsed: {t_end - t_start} seconds")
model_fs.write("./adata_cd8_zheng_models/cell_typist_model.pkl")
sc.pp.normalize_total(adata_cd8_zheng_test, target_sum=1e4)
sc.pp.log1p(adata_cd8_zheng_test)
predictions = celltypist.annotate(
adata_cd8_zheng_test, model="./adata_cd8_zheng_models/cell_typist_model.pkl"
)
col = list(predictions.probability_matrix.columns)
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(
list(
map(
lambda x: col.index(x),
adata_cd8_zheng_test.obs["cell_subtype_zheng_2021"],
)
)
).unsqueeze(1),
len(col),
)
from sklearn.metrics import roc_auc_score
roc_auc_scores_cell_typist.append(
roc_auc_score(y, predictions.probability_matrix.to_numpy(), average=None).mean()
)
Left one study for prediction¶
roc_auc_scores_scatlasvae_unseen = {}
for i in np.unique(adata_cd8_zheng.obs["study_name"]):
if i not in roc_auc_scores_scatlasvae_unseen.keys():
adata_cd8_zheng_train = adata_cd8_zheng[adata_cd8_zheng.obs["study_name"] != i]
adata_cd8_zheng_test = adata_cd8_zheng[adata_cd8_zheng.obs["study_name"] == i]
adata_cd8_zheng_test.obs["cell_subtype_zheng_2021"] = pd.Categorical(
adata_cd8_zheng_test.obs["cell_subtype_zheng_2021"],
categories=adata_cd8_zheng_train.obs[
"cell_subtype_zheng_2021"
].cat.categories,
)
vae_model = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_zheng_train,
batch_key=["sample_name", "study_name"],
batch_embedding="embedding",
batch_hidden_dim=64,
label_key="cell_subtype_zheng_2021",
device="cuda:0",
)
vae_model.fit(max_epoch=10, lr=5e-5, pred_weight=5.0)
checkpoint_path = "./adata_cd8_zheng_models/left.model"
vae_model.save_to_disk(checkpoint_path)
# adata_cd8_zheng_test = scatlasvae.model.scAtlasVAE.setup_anndata(
# adata_cd8_zheng_test,
# checkpoint_path
# )
import torch
state_dict = torch.load(checkpoint_path, map_location="cpu")
config = state_dict["model_config"]
vae_model_test = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_zheng_test,
pretrained_state_dict=state_dict["model_state_dict"],
**config,
)
from sklearn.metrics import roc_auc_score
vae_model_test.eval()
label_predictions = vae_model_test.predict_labels().detach().cpu().numpy()
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(list(map(lambda x: x[-2], vae_model_test._dataset)))[
[vae_model_test._shuffle_indices]
].unsqueeze(1),
vae_model_test.n_label + 1,
)
indices = adata_cd8_zheng_test.obs["cell_subtype_zheng_2021"] != "undefined"
sel = y[indices][:, :-1].sum(0) > 0
roc_auc_scores_scatlasvae_unseen[i] = roc_auc_score(
y[indices][:, :-1][:, sel], label_predictions[indices][:, sel], average=None
).mean()
roc_auc_scores_scatlasvae_seen = {}
for i in np.unique(adata_cd8_zheng.obs["study_name"]):
if i not in roc_auc_scores_scatlasvae_seen.keys():
adata_cd8_zheng.obs["cell_subtype_zheng_2021_scANVI"] = list(
adata_cd8_zheng.obs["cell_subtype_zheng_2021"]
)
adata_cd8_zheng.obs.loc[
adata_cd8_zheng.obs["study_name"] == i, "cell_subtype_zheng_2021_scANVI"
] = "undefined"
vae_model = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_zheng,
batch_key=["sample_name", "study_name"],
batch_embedding="embedding",
batch_hidden_dim=64,
label_key="cell_subtype_zheng_2021_scANVI",
device="cuda:0",
)
vae_model.fit(max_epoch=10, lr=5e-5, pred_weight=5.0)
from sklearn.metrics import roc_auc_score
vae_model.eval()
col = list(vae_model.label_category.categories)
label_predictions = vae_model.predict_labels().detach().cpu().numpy()
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(
list(
map(
lambda x: col.index(x),
adata_cd8_zheng.obs["cell_subtype_zheng_2021"],
)
)
).unsqueeze(1),
vae_model.n_label + 1,
)
indices = np.array(
adata_cd8_zheng.obs["cell_subtype_zheng_2021"] != "undefined"
) & np.array(adata_cd8_zheng.obs["study_name"] == i)
sel = y[indices][:, :-1].sum(0) > 0
roc_auc_scores_scatlasvae_seen[i] = roc_auc_score(
y[indices][:, :-1][:, sel], label_predictions[indices][:, sel], average=None
).mean()
roc_auc_scores_scanvi = {}
for i in np.unique(adata_cd8_zheng.obs["study_name"]):
if i not in roc_auc_scores_scanvi.keys():
adata_cd8_zheng.obs["cell_subtype_zheng_2021_scANVI"] = list(
adata_cd8_zheng.obs["cell_subtype_zheng_2021"]
)
adata_cd8_zheng.obs.loc[
adata_cd8_zheng.obs["study_name"] == i, "cell_subtype_zheng_2021_scANVI"
] = "undefined"
scvi.model.SCANVI.setup_anndata(
adata_cd8_zheng,
batch_key="sample_name",
categorical_covariate_keys=["study_name"],
labels_key="cell_subtype_zheng_2021_scANVI",
unlabeled_category="undefined",
)
model = scvi.model.SCANVI(adata_cd8_zheng)
model.train(max_epochs=10)
scanvi_prediction = model.predict(soft=True)
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(
list(
map(
lambda x: col.index(x),
adata_cd8_zheng.obs["cell_subtype_zheng_2021"],
)
)
).unsqueeze(1),
len(col),
)
indices = np.array(
adata_cd8_zheng.obs["cell_subtype_zheng_2021"] != "undefined"
) & np.array(adata_cd8_zheng.obs["study_name"] == i)
labels = y[indices][:, :-1]
roc_auc_scores_scanvi[i] = roc_auc_score(
labels[:, labels.sum(0) > 0],
scanvi_prediction.to_numpy()[indices, :][:, labels.sum(0) > 0],
average=None,
).mean()
roc_auc_scores_scpoli_unseen = {}
for i in np.unique(adata_cd8_zheng.obs["study_name"]):
if i not in roc_auc_scores_scpoli_unseen.keys():
try:
adata_cd8_zheng_train = adata_cd8_zheng[
adata_cd8_zheng.obs["study_name"] != i
]
adata_cd8_zheng_test = adata_cd8_zheng[
adata_cd8_zheng.obs["study_name"] == i
]
scpoli_model = scarches.models.scpoli.scPoli(
adata=adata_cd8_zheng_train[
adata_cd8_zheng_train.obs["cell_subtype_zheng_2021"] != "undefined"
],
condition_keys=["study_name"],
cell_type_keys=["cell_subtype_zheng_2021"],
unknown_ct_names=["undefined"],
recon_loss="zinb",
)
early_stopping_kwargs = {
"early_stopping_metric": "val_prototype_loss",
"mode": "min",
"threshold": 0,
"patience": 20,
"reduce_lr": True,
"lr_patience": 13,
"lr_factor": 0.1,
}
scpoli_model.train(
n_epochs=10,
pretraining_epochs=5,
early_stopping_kwargs=early_stopping_kwargs,
eta=5,
prototype_training=True,
)
# adata_cd8_zheng_test.obs.pop('cell_subtype_zheng_2021')
# adata_cd8_zheng_test.obs['cell_subtype_zheng_2021'] = 'undefined'
scpoli_query = scarches.models.scpoli.scPoli.load_query_data(
adata=adata_cd8_zheng_test,
reference_model=scpoli_model,
labeled_indices=[],
)
scpoli_query.train(n_epochs=10, pretraining_epochs=5, eta=10)
c = list(scpoli_query.cell_types_.keys())
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(
list(
map(
lambda x: c.index(x) if x in c else len(c),
adata_cd8_zheng_test.obs["cell_subtype_zheng_2021"],
)
)
).unsqueeze(1),
len(c) + 1,
)
results_dict = scpoli_query.classify(
adata_cd8_zheng_test, scale_uncertainties=True
)
indices = adata_cd8_zheng_test.obs["cell_subtype_zheng_2021"] != "undefined"
from sklearn.metrics import roc_auc_score
sel = y[indices][:, :-1].sum(0) > 0
roc_auc_scores_scpoli_unseen[i] = roc_auc_score(
y[indices][:, :-1][:, sel],
1
- results_dict["cell_subtype_zheng_2021"]["weighted_distances"][:, :][
indices
][:, sel],
average=None,
).mean()
except:
continue
roc_auc_scores_celltypist_left_study = {}
for i in np.unique(adata_cd8_zheng.obs["study_name"]):
if i not in roc_auc_scores_celltypist_left_study.keys():
try:
adata_cd8_zheng_train = adata_cd8_zheng[
adata_cd8_zheng.obs["study_name"] != i
]
adata_cd8_zheng_test = adata_cd8_zheng[
adata_cd8_zheng.obs["study_name"] == i
]
adata_cd8_zheng_train = adata_cd8_zheng_train[
adata_cd8_zheng_train.obs["cell_subtype_zheng_2021"] != "undefined"
]
adata_cd8_zheng_test = adata_cd8_zheng_test[
adata_cd8_zheng_test.obs["cell_subtype_zheng_2021"] != "undefined"
]
sc.pp.normalize_total(adata_cd8_zheng_train, target_sum=1e4)
sc.pp.log1p(adata_cd8_zheng_train)
import time
import celltypist
t_start = time.time()
model_fs = celltypist.train(
adata_cd8_zheng_train,
"cell_subtype_zheng_2021",
n_jobs=96,
max_iter=5,
use_SGD=True,
)
t_end = time.time()
print(f"Time elapsed: {t_end - t_start} seconds")
model_fs.write("./adata_cd8_zheng_models/cell_typist_model.pkl")
sc.pp.normalize_total(adata_cd8_zheng_test, target_sum=1e4)
sc.pp.log1p(adata_cd8_zheng_test)
predictions = celltypist.annotate(
adata_cd8_zheng_test,
model="./adata_cd8_zheng_models/cell_typist_model.pkl",
)
col = list(predictions.probability_matrix.columns)
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(
list(
map(
lambda x: col.index(x),
adata_cd8_zheng_test.obs["cell_subtype_zheng_2021"],
)
)
).unsqueeze(1),
len(col),
)
sel = y.sum(0) > 0
from sklearn.metrics import roc_auc_score
roc_auc_scores_celltypist_left_study[i] = roc_auc_score(
y[:, sel],
predictions.probability_matrix.to_numpy()[:, sel],
average=None,
).mean()
except:
continue
Benchmark Chu et al., 2023 Dataset¶
Benchmarking integration performance¶
##############
# scAtlasVAE #
##############
vae_model = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_chu,
batch_key="study_name",
batch_embedding="embedding",
batch_hidden_dim=64,
device="cuda:0",
)
vae_model.fit(max_epoch=10, lr=5e-5)
adata_cd8_chu.obsm["X_gex"] = vae_model.get_latent_embedding(show_progress=True)
vae_model = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_chu,
batch_key="study_name",
batch_embedding="embedding",
batch_hidden_dim=64,
label_key="cell_subtype_chu_2023",
device="cuda:0",
)
vae_model.fit(max_epoch=10, lr=5e-5)
adata_cd8_chu.obsm["X_gex_supervised"] = vae_model.get_latent_embedding(
show_progress=True
)
###################
# scVI and scANVI #
###################
scvi.model.SCVI.setup_anndata(adata_cd8_chu, batch_key="study_name")
model = scvi.model.SCVI(adata_cd8_chu)
model.train(max_epochs=10)
adata_cd8_chu.obsm["X_scVI"] = model.get_latent_representation()
scvi.model.SCANVI.setup_anndata(
adata_cd8_chu,
batch_key="study_name",
labels_key="cell_subtype_chu_2023",
unlabeled_category="undefined",
)
model = scvi.model.SCANVI(adata_cd8_chu)
model.train(max_epochs=10)
adata_cd8_chu.obsm["X_scANVI"] = model.get_latent_representation()
##########
# scPoli #
##########
scpoli_model = scarches.models.scpoli.scPoli(
adata=adata_cd8_chu,
condition_keys=["study_name"],
recon_loss="zinb",
)
early_stopping_kwargs = {
"early_stopping_metric": "val_prototype_loss",
"mode": "min",
"threshold": 0,
"patience": 20,
"reduce_lr": True,
"lr_patience": 13,
"lr_factor": 0.1,
}
scpoli_model.train(
n_epochs=10,
pretraining_epochs=5,
early_stopping_kwargs=early_stopping_kwargs,
eta=5,
)
for i in tqdm.trange(0, len(adata_cd8_chu), 320):
Z.append(scpoli_model.get_latent(adata_cd8_chu[i : i + 320]))
adata_cd8_chu.obsm["X_scPoli"] = Z
scpoli_model = scarches.models.scpoli.scPoli(
adata=adata_cd8_chu,
condition_keys=["study_name"],
recon_loss="zinb",
cell_type_keys=["cell_subtype_chu_2023"],
unknown_ct_names=["undefined"],
)
scpoli_model.train(
n_epochs=10,
pretraining_epochs=5,
early_stopping_kwargs=early_stopping_kwargs,
eta=5,
)
Z = []
import tqdm
# scpoli_model.
for i in tqdm.trange(0, len(adata_cd8_chu), 320):
Z.append(scpoli_model.get_latent(adata_cd8_chu[i : i + 320]))
adata_cd8_chu.obsm["X_scPoli_supervised"] = Z
##########
# SCALEX #
##########
os.chdir("~/Biosoft/SCALEX")
from scalex import SCALEX
adata_cd8_chu.obsm["X"] = adata_cd8_chu.X
adata_cd8_chu_scalex = SCALEX(
adata_cd8_chu,
batch_name=["study_name"],
n_top_features=adata_cd8_chu.shape[1],
min_cells=1,
min_features=1,
use_layer="X",
ignore_umap=True,
)
adata_cd8_chu.obsm["X_SCALEX"] = adata_cd8_chu_scalex.obsm["latent"]
###########
# Harmony #
###########
from harmony import harmonize
sc.pp.normalize_total(adata_cd8_chu)
sc.pp.log1p(adata_cd8_chu)
sc.tl.pca(adata_cd8_chu)
adata_cd8_chu.obsm["X_harmony"] = harmonize(
adata_cd8_chu.obsm["X_pca"], adata_cd8_chu.obs, batch_key="study_name"
)
#############
# Scanoarma #
#############
def scanoramaCorrectMerge(adata, groupby="study_name"):
adatas = []
group_names = list(np.unique(adata.obs[groupby]))
for i in group_names:
adatas.append(adata[adata.obs[groupby] == i])
corrected = scanorama.correct_scanpy(adatas, dimred=10, return_dimred=True)
return sc.concat(corrected)[adata.obs.index]
adata_cd8_chu.obsm["X_scanorama"] = scanoramaCorrectMerge(
adata_cd8_chu, groupby="study_name"
).obsm["X_scanorama"]
Benchmarking label transfer performance¶
Left 5% random cells from study¶
import sklearn
from sklearn.metrics import roc_auc_score
roc_auc_scores_scatlasvae_zero_shot = {
i: [] for i in [1.0, 2.0, 3.0, 4.0, 5.0, 6.0, 7.0]
}
for i in [1.0, 2.0, 3.0, 4.0, 5.0, 6.0, 7.0]:
for _ in range(3):
l = list(range(adata_cd8_chu.shape[0]))
train, test = sklearn.model_selection.train_test_split(l, test_size=0.05)
adata_cd8_chu_train = adata_cd8_chu[train]
adata_cd8_chu_test = adata_cd8_chu[test]
vae_model = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_chu_train,
batch_key=["study_name"],
batch_embedding="embedding",
batch_hidden_dim=64,
label_key="cell_subtype_chu_2023",
device="cuda:0",
)
vae_model.fit(
max_epoch=10,
lr=5e-5,
pred_weight=i,
)
checkpoint_path = "./adata_cd8_chu_models/left.model"
vae_model.save_to_disk(checkpoint_path)
import torch
state_dict = torch.load(checkpoint_path, map_location="cpu")
config = state_dict["model_config"]
vae_model_test = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_chu_test,
pretrained_state_dict=state_dict["model_state_dict"],
**config,
)
from sklearn.metrics import roc_auc_score
vae_model_test.eval()
label_predictions = vae_model_test.predict_labels().detach().cpu().numpy()
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(list(map(lambda x: x[-2], vae_model_test._dataset)))[
[vae_model_test._shuffle_indices]
].unsqueeze(1),
vae_model_test.n_label + 1,
)
roc_auc_scores_scatlasvae_zero_shot[i].append(
roc_auc_score(y, label_predictions, average=None).mean()
)
roc_auc_scores_scatlasvae_full_shot = {
i: [] for i in [1.0, 2.0, 3.0, 4.0, 5.0, 6.0, 7.0]
}
for i in [1.0, 2.0, 3.0, 4.0, 5.0, 6.0, 7.0]:
for _ in range(10):
l = list(range(adata_cd8_chu.shape[0]))
train, test = sklearn.model_selection.train_test_split(l, test_size=0.05)
adata_cd8_chu.obs["cell_subtype_chu_2023_scAtlasVAE"] = list(
adata_cd8_chu.obs["cell_subtype_chu_2023"]
)
adata_cd8_chu.obs.iloc[
test,
list(adata_cd8_chu.obs.columns).index("cell_subtype_chu_2023_scAtlasVAE"),
] = "undefined"
vae_model = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_chu,
batch_key=["study_name"],
batch_embedding="embedding",
batch_hidden_dim=64,
label_key="cell_subtype_chu_2023_scANVI",
device="cuda:0",
)
vae_model.fit(max_epoch=10, lr=5e-5, pred_weight=i)
from sklearn.metrics import roc_auc_score
vae_model.eval()
label_predictions = vae_model.predict_labels().detach().cpu().numpy()
col = list(vae_model.label_category.categories)
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(
list(
map(
lambda x: col.index(x),
adata_cd8_chu.obs["cell_subtype_chu_2023"],
)
)
).unsqueeze(1),
vae_model.n_label + 1,
)
test_set = set(test)
indices = np.array(
adata_cd8_chu.obs["cell_subtype_chu_2023"] != "undefined"
) & np.array(list(map(lambda x: x in test_set, range(len(adata_cd8_chu)))))
sel = y[indices][:, :-1].sum(0) > 0
roc_auc_scores_scatlasvae_full_shot[i].append(
roc_auc_score(
y[indices][:, :-1][:, sel],
label_predictions[indices][:, sel],
average=None,
).mean()
)
scANVI_roc_auc_scores = []
for _ in range(10):
l = list(range(adata_cd8_chu.shape[0]))
train, test = sklearn.model_selection.train_test_split(l, test_size=0.05)
adata_cd8_chu.obs["cell_subtype_chu_2023_scANVI"] = list(
adata_cd8_chu.obs["cell_subtype_chu_2023"]
)
adata_cd8_chu.obs.iloc[
test, list(adata_cd8_chu.obs.columns).index("cell_subtype_chu_2023_scANVI")
] = "undefined"
scvi.model.SCANVI.setup_anndata(
adata_cd8_chu,
batch_key="study_name",
labels_key="cell_subtype_chu_2023_scANVI",
unlabeled_category="undefined",
)
model = scvi.model.SCANVI(adata_cd8_chu)
model.train(max_epochs=10)
scanvi_prediction = model.predict(soft=True)
col = list(scanvi_prediction.columns) + ["undefined"]
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(
list(
map(lambda x: col.index(x), adata_cd8_chu.obs["cell_subtype_chu_2023"])
)
).unsqueeze(1),
len(col),
)
test = set(test)
indices = list(
map(
lambda x: x[1] != "undefined" and x[0] in test,
enumerate(adata_cd8_chu.obs["cell_subtype_chu_2023"]),
)
)
scANVI_roc_auc_scores.append(
roc_auc_score(y[indices], scanvi_prediction.to_numpy()[indices])
)
scpoli_roc_auc_scores = []
for _ in range(10):
l = list(range(adata_cd8_chu.shape[0]))
train, test = sklearn.model_selection.train_test_split(l, test_size=0.05)
adata_cd8_chu_train = adata_cd8_chu[train]
adata_cd8_chu_test = adata_cd8_chu[test]
scpoli_model = scarches.models.scpoli.scPoli(
adata=adata_cd8_chu_train[
adata_cd8_chu_train.obs["cell_subtype_chu_2023"] != "undefined"
],
condition_keys=["study_name"],
cell_type_keys=["cell_subtype_chu_2023"],
unknown_ct_names=["undefined"],
recon_loss="zinb",
)
early_stopping_kwargs = {
"early_stopping_metric": "val_prototype_loss",
"mode": "min",
"threshold": 0,
"patience": 20,
"reduce_lr": True,
"lr_patience": 13,
"lr_factor": 0.1,
}
scpoli_model.train(
n_epochs=10,
pretraining_epochs=5,
early_stopping_kwargs=early_stopping_kwargs,
eta=5,
prototype_training=True,
)
# adata_cd8_chu_test.obs.pop('cell_subtype_chu_2023')
# adata_cd8_chu_test.obs['cell_subtype_chu_2023'] = 'undefined'
scpoli_query = scarches.models.scpoli.scPoli.load_query_data(
adata=adata_cd8_chu_test,
reference_model=scpoli_model,
labeled_indices=[],
)
scpoli_query.train(n_epochs=10, pretraining_epochs=5, eta=10)
c = list(scpoli_query.cell_types_.keys())
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(
list(
map(
lambda x: c.index(x) if x in c else vae_model_test.n_label,
adata_cd8_chu_test.obs["cell_subtype_chu_2023"],
)
)
).unsqueeze(1),
len(c),
)
results_dict = scpoli_query.classify(adata_cd8_chu_test, scale_uncertainties=True)
indices = adata_cd8_chu_test.obs["cell_subtype_chu_2023"] != "undefined"
scpoli_roc_auc_scores.append(
roc_auc_score(
y[indices],
1 - results_dict["cell_subtype_chu_2023"]["weighted_distances"][indices],
)
)
roc_auc_scores_cell_typist = []
for _ in range(10):
l = list(range(adata_cd8_chu.shape[0]))
train, test = sklearn.model_selection.train_test_split(l, test_size=0.05)
adata_cd8_chu_train = adata_cd8_chu[train]
adata_cd8_chu_train = adata_cd8_chu_train[
adata_cd8_chu_train.obs["cell_subtype_chu_2023"] != "undefined"
]
adata_cd8_chu_test = adata_cd8_chu[test]
adata_cd8_chu_test = adata_cd8_chu_test[
adata_cd8_chu_test.obs["cell_subtype_chu_2023"] != "undefined"
]
sc.pp.normalize_total(adata_cd8_chu_train, target_sum=1e4)
sc.pp.log1p(adata_cd8_chu_train)
t_start = time.time()
model_fs = celltypist.train(
adata_cd8_chu_train,
"cell_subtype_chu_2023",
n_jobs=96,
max_iter=5,
use_SGD=True,
)
t_end = time.time()
print(f"Time elapsed: {t_end - t_start} seconds")
model_fs.write("./adata_cd8_chu_models/cell_typist_model.pkl")
sc.pp.normalize_total(adata_cd8_chu_test, target_sum=1e4)
sc.pp.log1p(adata_cd8_chu_test)
predictions = celltypist.annotate(
adata_cd8_chu_test, model="./adata_cd8_chu_models/cell_typist_model.pkl"
)
col = list(predictions.probability_matrix.columns)
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(
list(
map(
lambda x: col.index(x),
adata_cd8_chu_test.obs["cell_subtype_chu_2023"],
)
)
).unsqueeze(1),
len(col),
)
from sklearn.metrics import roc_auc_score
roc_auc_scores_cell_typist.append(
roc_auc_score(y, predictions.probability_matrix.to_numpy(), average=None).mean()
)
Left one study for prediction¶
roc_auc_scores_scatlasvae_unseen = {}
for i in np.unique(adata_cd8_chu.obs["study_name"]):
if i not in roc_auc_scores_scatlasvae_unseen.keys():
adata_cd8_chu_train = adata_cd8_chu[adata_cd8_chu.obs["study_name"] != i]
adata_cd8_chu_test = adata_cd8_chu[adata_cd8_chu.obs["study_name"] == i]
adata_cd8_chu_test.obs["cell_subtype_chu_2023"] = pd.Categorical(
adata_cd8_chu_test.obs["cell_subtype_chu_2023"],
categories=adata_cd8_chu_train.obs["cell_subtype_chu_2023"].cat.categories,
)
vae_model = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_chu_train,
batch_key=["study_name"],
batch_embedding="embedding",
batch_hidden_dim=64,
label_key="cell_subtype_chu_2023",
device="cuda:0",
)
vae_model.fit(max_epoch=10, lr=5e-5, pred_weight=5.0)
checkpoint_path = "./adata_cd8_chu_models/left.model"
vae_model.save_to_disk(checkpoint_path)
# adata_cd8_chu_test = scatlasvae.model.scAtlasVAE.setup_anndata(
# adata_cd8_chu_test,
# checkpoint_path
# )
import torch
state_dict = torch.load(checkpoint_path, map_location="cpu")
config = state_dict["model_config"]
vae_model_test = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_chu_test,
pretrained_state_dict=state_dict["model_state_dict"],
**config,
)
from sklearn.metrics import roc_auc_score
vae_model_test.eval()
label_predictions = vae_model_test.predict_labels().detach().cpu().numpy()
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(list(map(lambda x: x[-2], vae_model_test._dataset)))[
[vae_model_test._shuffle_indices]
].unsqueeze(1),
vae_model_test.n_label + 1,
)
indices = adata_cd8_chu_test.obs["cell_subtype_chu_2023"] != "undefined"
sel = y[indices][:, :-1].sum(0) > 0
roc_auc_scores_scatlasvae_unseen[i] = roc_auc_score(
y[indices][:, :-1][:, sel], label_predictions[indices][:, sel], average=None
).mean()
roc_auc_scores_scatlasvae_seen = {}
for i in np.unique(adata_cd8_chu.obs["study_name"]):
if i not in roc_auc_scores_scatlasvae_seen.keys():
adata_cd8_chu.obs["cell_subtype_chu_2023_scANVI"] = list(
adata_cd8_chu.obs["cell_subtype_chu_2023"]
)
adata_cd8_chu.obs.loc[
adata_cd8_chu.obs["study_name"] == i, "cell_subtype_chu_2023_scANVI"
] = "undefined"
vae_model = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_chu,
batch_key=["study_name"],
batch_embedding="embedding",
batch_hidden_dim=64,
label_key="cell_subtype_chu_2023_scANVI",
device="cuda:0",
)
vae_model.fit(max_epoch=10, lr=5e-5, pred_weight=5.0)
from sklearn.metrics import roc_auc_score
vae_model.eval()
col = list(vae_model.label_category.categories)
label_predictions = vae_model.predict_labels().detach().cpu().numpy()
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(
list(
map(
lambda x: col.index(x),
adata_cd8_chu.obs["cell_subtype_chu_2023"],
)
)
).unsqueeze(1),
vae_model.n_label + 1,
)
indices = np.array(
adata_cd8_chu.obs["cell_subtype_chu_2023"] != "undefined"
) & np.array(adata_cd8_chu.obs["study_name"] == i)
sel = y[indices][:, :-1].sum(0) > 0
roc_auc_scores_scatlasvae_seen[i] = roc_auc_score(
y[indices][:, :-1][:, sel], label_predictions[indices][:, sel], average=None
).mean()
roc_auc_scores_scanvi = {}
for i in np.unique(adata_cd8_chu.obs["study_name"]):
if i not in roc_auc_scores_scanvi.keys():
adata_cd8_chu.obs["cell_subtype_chu_2023_scANVI"] = list(
adata_cd8_chu.obs["cell_subtype_chu_2023"]
)
adata_cd8_chu.obs.loc[
adata_cd8_chu.obs["study_name"] == i, "cell_subtype_chu_2023_scANVI"
] = "undefined"
scvi.model.SCANVI.setup_anndata(
adata_cd8_chu,
batch_key="study_name",
labels_key="cell_subtype_chu_2023_scANVI",
unlabeled_category="undefined",
)
model = scvi.model.SCANVI(adata_cd8_chu)
model.train(max_epochs=10)
scanvi_prediction = model.predict(soft=True)
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(
list(
map(
lambda x: col.index(x),
adata_cd8_chu.obs["cell_subtype_chu_2023"],
)
)
).unsqueeze(1),
len(col),
)
indices = np.array(
adata_cd8_chu.obs["cell_subtype_chu_2023"] != "undefined"
) & np.array(adata_cd8_chu.obs["study_name"] == i)
labels = y[indices][:, :-1]
roc_auc_scores_scanvi[i] = roc_auc_score(
labels[:, labels.sum(0) > 0],
scanvi_prediction.to_numpy()[indices, :][:, labels.sum(0) > 0],
average=None,
).mean()
roc_auc_scores_scpoli_unseen = {}
for i in np.unique(adata_cd8_chu.obs["study_name"]):
if i not in roc_auc_scores_scpoli_unseen.keys():
try:
adata_cd8_chu_train = adata_cd8_chu[adata_cd8_chu.obs["study_name"] != i]
adata_cd8_chu_test = adata_cd8_chu[adata_cd8_chu.obs["study_name"] == i]
scpoli_model = scarches.models.scpoli.scPoli(
adata=adata_cd8_chu_train[
adata_cd8_chu_train.obs["cell_subtype_chu_2023"] != "undefined"
],
condition_keys=["study_name"],
cell_type_keys=["cell_subtype_chu_2023"],
unknown_ct_names=["undefined"],
recon_loss="zinb",
)
early_stopping_kwargs = {
"early_stopping_metric": "val_prototype_loss",
"mode": "min",
"threshold": 0,
"patience": 20,
"reduce_lr": True,
"lr_patience": 13,
"lr_factor": 0.1,
}
scpoli_model.train(
n_epochs=10,
pretraining_epochs=5,
early_stopping_kwargs=early_stopping_kwargs,
eta=5,
prototype_training=True,
)
# adata_cd8_chu_test.obs.pop('cell_subtype_chu_2023')
# adata_cd8_chu_test.obs['cell_subtype_chu_2023'] = 'undefined'
scpoli_query = scarches.models.scpoli.scPoli.load_query_data(
adata=adata_cd8_chu_test,
reference_model=scpoli_model,
labeled_indices=[],
)
scpoli_query.train(n_epochs=10, pretraining_epochs=5, eta=10)
c = list(scpoli_query.cell_types_.keys())
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(
list(
map(
lambda x: c.index(x) if x in c else len(c),
adata_cd8_chu_test.obs["cell_subtype_chu_2023"],
)
)
).unsqueeze(1),
len(c) + 1,
)
results_dict = scpoli_query.classify(
adata_cd8_chu_test, scale_uncertainties=True
)
indices = adata_cd8_chu_test.obs["cell_subtype_chu_2023"] != "undefined"
from sklearn.metrics import roc_auc_score
sel = y[indices][:, :-1].sum(0) > 0
roc_auc_scores_scpoli_unseen[i] = roc_auc_score(
y[indices][:, :-1][:, sel],
1
- results_dict["cell_subtype_chu_2023"]["weighted_distances"][:, :][
indices
][:, sel],
average=None,
).mean()
except:
continue
roc_auc_scores_celltypist_left_study = {}
for i in np.unique(adata_cd8_chu.obs["study_name"]):
if i not in roc_auc_scores_celltypist_left_study.keys():
try:
adata_cd8_chu_train = adata_cd8_chu[adata_cd8_chu.obs["study_name"] != i]
adata_cd8_chu_test = adata_cd8_chu[adata_cd8_chu.obs["study_name"] == i]
adata_cd8_chu_train = adata_cd8_chu_train[
adata_cd8_chu_train.obs["cell_subtype_chu_2023"] != "undefined"
]
adata_cd8_chu_test = adata_cd8_chu_test[
adata_cd8_chu_test.obs["cell_subtype_chu_2023"] != "undefined"
]
sc.pp.normalize_total(adata_cd8_chu_train, target_sum=1e4)
sc.pp.log1p(adata_cd8_chu_train)
import time
import celltypist
t_start = time.time()
model_fs = celltypist.train(
adata_cd8_chu_train,
"cell_subtype_chu_2023",
n_jobs=96,
max_iter=5,
use_SGD=True,
)
t_end = time.time()
print(f"Time elapsed: {t_end - t_start} seconds")
model_fs.write("./adata_cd8_chu_models/cell_typist_model.pkl")
sc.pp.normalize_total(adata_cd8_chu_test, target_sum=1e4)
sc.pp.log1p(adata_cd8_chu_test)
predictions = celltypist.annotate(
adata_cd8_chu_test, model="./adata_cd8_chu_models/cell_typist_model.pkl"
)
col = list(predictions.probability_matrix.columns)
y = scatlasvae.utils._tensor_utils.one_hot(
torch.tensor(
list(
map(
lambda x: col.index(x),
adata_cd8_chu_test.obs["cell_subtype_chu_2023"],
)
)
).unsqueeze(1),
len(col),
)
sel = y.sum(0) > 0
from sklearn.metrics import roc_auc_score
roc_auc_scores_celltypist_left_study[i] = roc_auc_score(
y[:, sel],
predictions.probability_matrix.to_numpy()[:, sel],
average=None,
).mean()
except:
continue
Benchmarking multi-atlas integration¶
vae_model = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_multi_atlas,
batch_key=['sample_name','study_name','atlas_name'],
label_key=['cell_subtype_3','cell_subtype_zheng_2021','cell_subtype_chu_2023']
batch_embedding='embedding',
batch_hidden_dim=64,
device='cuda:0'
)
vae_model.fit(max_epoch=10, lr=5e-5)
adata_cd8_multi_atlas.obsm["X_gex"] = vae_model.get_latent_representation()
scvi.model.SCVI.setup_anndata(
adata_cd8_multi_atlas,
batch_key='sample_name',
categorical_covariate_keys=['study_name','atlas_name'],
)
model = scvi.model.SCVI(adata_cd8_multi_atlas)
model.train(max_epochs=10)
adata_cd8_multi_atlas.obsm['X_scVI'] = model.get_latent_representation()
adata_cd8_multi_atlas.obs['merged_label'] = list(map(lambda x:list(filter(lambda z: z != 'undefined', x))[0], adata_cd8_multi_atlas.obs.loc[:,['cell_subtype_3','cell_subtype_zheng_2021','cell_subtype_chu_2023']].to_numpy()))
scvi.model.SCANVI.setup_anndata(
adata_cd8_multi_atlas,
batch_key='sample_name',
categorical_covariate_keys=['study_name','atlas_name'],
labels_key='merged_label',
unlabeled_category='undefined'
)
model = scvi.model.SCANVI(adata_cd8_multi_atlas)
model.train(max_epochs=10)
adata_cd8_multi_atlas.obsm['X_scANVI'] = model.get_latent_representation()
scpoli_model = scarches.models.scpoli.scPoli(
adata=adata_cd8_multi_atlas,
condition_keys=['sample_name', 'study_name'],
recon_loss='zinb',
cell_type_keys=['cell_subtype_3','cell_subtype_zheng_2021','cell_subtype_chu_2023'],
unknown_ct_names=['undefined']
)
early_stopping_kwargs = {
'early_stopping_metric': 'val_prototype_loss',
'mode': 'min',
'threshold': 0,
'patience': 20,
'reduce_lr': True,
'lr_patience': 13,
'lr_factor': 0.1,
}
scpoli_model.train(
n_epochs=50,
pretraining_epochs=40,
early_stopping_kwargs=early_stopping_kwargs,
eta=5,
)
Z = []
import tqdm
# scpoli_model.
for i in tqdm.trange(0,len(adata_cd8_multi_atlas),320):
Z.append(scpoli_model.get_latent(adata_cd8_multi_atlas[i:i+320]))
import numpy as np
Z = np.vstack(Z)
adata_cd8_multi_atlas.obsm['X_scPoli_supervised'] = Z
scpoli_model = scarches.models.scpoli.scPoli(
adata=adata_cd8_multi_atlas,
condition_keys=['sample_name', 'study_name'],
recon_loss='zinb',
# cell_type_keys=['cell_subtype_3','cell_subtype_zheng_2021','cell_subtype_chu_2023'],
unknown_ct_names=['undefined']
)
early_stopping_kwargs = {
'early_stopping_metric': 'val_prototype_loss',
'mode': 'min',
'threshold': 0,
'patience': 20,
'reduce_lr': True,
'lr_patience': 13,
'lr_factor': 0.1,
}
scpoli_model.train(
n_epochs=50,
pretraining_epochs=40,
early_stopping_kwargs=early_stopping_kwargs,
eta=5,
)
Z = []
import tqdm
# scpoli_model.
for i in tqdm.trange(0,len(adata_cd8_multi_atlas),320):
Z.append(scpoli_model.get_latent(adata_cd8_multi_atlas[i:i+320]))
import numpy as np
Z = np.vstack(Z)
adata_cd8_multi_atlas.obsm['X_scPoli'] = Z
os.chdir('/slurm/home/yrd/liulab/xueziwei/Biosoft/SCALEX')
from scalex import SCALEX
adata_cd8_multi_atlas.obsm['X'] = adata_cd8_multi_atlas.X
adata_cd8_multi_atlas_scalex = SCALEX(
adata_cd8_multi_atlas,
batch_name=['study_name'],
n_top_features=adata_cd8_multi_atlas.shape[1],
min_cells=1,
min_features=1,
use_layer='X',
ignore_umap=True
)
adata_cd8_multi_atlas.obsm['X_SCALEX'] = adata_cd8_multi_atlas_scalex.obsm['latent']
multi_atlas_silhouette_batch_result = {
"cell_subtype_revision": {
"sample_name": {},
"study_name": {},
},
"cell_subtype_zheng_2021": {
"sample_name": {},
"study_name": {},
},
"cell_subtype_chu_2023": {
"study_name": {},
},
}
for emb_key in ["X_gex", "X_scANVI", "X_scVI", "X_scPoli_supervised", "X_SCALEX"]:
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_revision"].isin(["undefined", "???"])
]
sc.pp.neighbors(adata_, use_rep=emb_key)
multi_atlas_silhouette_batch_result["cell_subtype_revision"]["sample_name"][
emb_key
] = scib.me.silhouette_batch(
adata_,
label_key="cell_subtype_revision",
batch_key="sample_name",
embed=emb_key,
)
multi_atlas_silhouette_batch_result["cell_subtype_revision"]["study_name"][
emb_key
] = scib.me.silhouette_batch(
adata_, label_key="cell_subtype_revision", batch_key="study_name", embed=emb_key
)
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_zheng_2021"].isin(["undefined"])
]
sc.pp.neighbors(adata_, use_rep=emb_key)
multi_atlas_silhouette_batch_result["cell_subtype_zheng_2021"]["sample_name"][
emb_key
] = scib.me.silhouette_batch(
adata_,
label_key="cell_subtype_zheng_2021",
batch_key="sample_name",
embed=emb_key,
)
multi_atlas_silhouette_batch_result["cell_subtype_zheng_2021"]["study_name"][
emb_key
] = scib.me.silhouette_batch(
adata_,
label_key="cell_subtype_zheng_2021",
batch_key="study_name",
embed=emb_key,
)
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_chu_2023"].isin(["undefined"])
]
sc.pp.neighbors(adata_, use_rep=emb_key)
multi_atlas_silhouette_batch_result["cell_subtype_chu_2023"]["study_name"][
emb_key
] = scib.me.silhouette_batch(
adata_, label_key="cell_subtype_chu_2023", batch_key="study_name", embed=emb_key
)
multi_atlas_silhouette_result = {
"cell_subtype_revision": {
"sample_name": {},
"study_name": {},
},
"cell_subtype_zheng_2021": {
"sample_name": {},
"study_name": {},
},
"cell_subtype_chu_2023": {
"study_name": {},
},
}
for emb_key in ["X_gex", "X_scANVI", "X_scVI", "X_scPoli_supervised", "X_SCALEX"]:
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_revision"].isin(["undefined", "???"])
]
sc.pp.neighbors(adata_, use_rep=emb_key)
multi_atlas_silhouette_result["cell_subtype_revision"]["sample_name"][
emb_key
] = scib.me.silhouette(adata_, label_key="cell_subtype_revision", embed=emb_key)
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_zheng_2021"].isin(["undefined"])
]
sc.pp.neighbors(adata_, use_rep=emb_key)
multi_atlas_silhouette_result["cell_subtype_zheng_2021"]["study_name"][
emb_key
] = scib.me.silhouette(adata_, label_key="cell_subtype_zheng_2021", embed=emb_key)
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_chu_2023"].isin(["undefined"])
]
sc.pp.neighbors(adata_, use_rep=emb_key)
multi_atlas_silhouette_result["cell_subtype_chu_2023"]["study_name"][
emb_key
] = scib.me.silhouette(adata_, label_key="cell_subtype_chu_2023", embed=emb_key)
multi_atlas_graph_connectivity_result = {
"cell_subtype_revision": {},
"cell_subtype_zheng_2021": {},
"cell_subtype_chu_2023": {},
}
for emb_key in ["X_gex", "X_scANVI", "X_scVI", "X_scPoli_supervised", "X_SCALEX"]:
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_revision"].isin(["undefined", "???"])
]
sc.pp.neighbors(adata_, use_rep=emb_key)
multi_atlas_graph_connectivity_result["cell_subtype_revision"][
emb_key
] = scib.me.graph_connectivity(adata_, "cell_subtype_revision")
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_zheng_2021"].isin(["undefined"])
]
sc.pp.neighbors(adata_, use_rep=emb_key)
multi_atlas_graph_connectivity_result["cell_subtype_zheng_2021"][
emb_key
] = scib.me.graph_connectivity(
adata_,
label_key="cell_subtype_zheng_2021",
)
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_chu_2023"].isin(["undefined"])
]
sc.pp.neighbors(adata_, use_rep=emb_key)
multi_atlas_graph_connectivity_result["cell_subtype_chu_2023"][
emb_key
] = scib.me.graph_connectivity(
adata_,
label_key="cell_subtype_chu_2023",
)
multi_atlas_pcr_comparison_result = {
"cell_subtype_revision": {},
"cell_subtype_zheng_2021": {},
"cell_subtype_chu_2023": {},
}
for emb_key in ["X_gex", "X_scANVI", "X_scVI", "X_scPoli_supervised", "X_SCALEX"]:
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_revision"].isin(["undefined"])
]
multi_atlas_pcr_comparison_result["cell_subtype_revision"][
emb_key
] = scib.me.pcr_comparison(
adata_, adata_, covariate="study_name", embed=emb_key, n_comps=50
)
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_zheng_2021"].isin(["undefined"])
]
multi_atlas_pcr_comparison_result["cell_subtype_zheng_2021"][
emb_key
] = scib.me.pcr_comparison(
adata_, adata_, covariate="study_name", embed=emb_key, n_comps=30
)
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_chu_2023"].isin(["undefined"])
]
multi_atlas_pcr_comparison_result["cell_subtype_chu_2023"][
emb_key
] = scib.me.pcr_comparison(
adata_, adata_, covariate="study_name", embed=emb_key, n_comps=30
)
multi_atlas_isolated_labels_asw_result = {
"cell_subtype_revision": {
"sample_name": {},
"study_name": {},
},
"cell_subtype_zheng_2021": {
"sample_name": {},
"study_name": {},
},
"cell_subtype_chu_2023": {
"study_name": {},
},
}
for emb_key in ["X_gex", "X_scANVI", "X_scVI", "X_scPoli_supervised", "X_SCALEX"]:
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_revision"].isin(["undefined", "???"])
]
sc.pp.neighbors(adata_, use_rep=emb_key)
multi_atlas_isolated_labels_asw_result["cell_subtype_revision"]["sample_name"][
emb_key
] = scib.me.isolated_labels_asw(
adata_,
label_key="cell_subtype_revision",
batch_key="sample_name",
embed=emb_key,
)
multi_atlas_isolated_labels_asw_result["cell_subtype_revision"]["study_name"][
emb_key
] = scib.me.isolated_labels_asw(
adata_, label_key="cell_subtype_revision", batch_key="study_name", embed=emb_key
)
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_zheng_2021"].isin(["undefined"])
]
sc.pp.neighbors(adata_, use_rep=emb_key)
multi_atlas_isolated_labels_asw_result["cell_subtype_zheng_2021"]["sample_name"][
emb_key
] = scib.me.isolated_labels_asw(
adata_,
label_key="cell_subtype_zheng_2021",
batch_key="sample_name",
embed=emb_key,
)
multi_atlas_isolated_labels_asw_result["cell_subtype_zheng_2021"]["study_name"][
emb_key
] = scib.me.isolated_labels_asw(
adata_,
label_key="cell_subtype_zheng_2021",
batch_key="study_name",
embed=emb_key,
)
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_chu_2023"].isin(["undefined"])
]
sc.pp.neighbors(adata_, use_rep=emb_key)
multi_atlas_isolated_labels_asw_result["cell_subtype_chu_2023"]["study_name"][
emb_key
] = scib.me.isolated_labels_asw(
adata_, label_key="cell_subtype_chu_2023", batch_key="study_name", embed=emb_key
)
multi_atlas_isolated_labels_f1_result = {
"cell_subtype_revision": {
"sample_name": {},
"study_name": {},
},
"cell_subtype_zheng_2021": {
"sample_name": {},
"study_name": {},
},
"cell_subtype_chu_2023": {
"study_name": {},
},
}
for emb_key in ["X_gex", "X_scANVI", "X_scVI", "X_scPoli_supervised", "X_SCALEX"]:
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_revision"].isin(["undefined", "???"])
]
sc.pp.neighbors(adata_, use_rep=emb_key)
multi_atlas_isolated_labels_f1_result["cell_subtype_revision"]["sample_name"][
emb_key
] = scib.me.isolated_labels_f1(
adata_,
label_key="cell_subtype_revision",
batch_key="sample_name",
embed=emb_key,
)
multi_atlas_isolated_labels_f1_result["cell_subtype_revision"]["study_name"][
emb_key
] = scib.me.isolated_labels_f1(
adata_, label_key="cell_subtype_revision", batch_key="study_name", embed=emb_key
)
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_zheng_2021"].isin(["undefined"])
]
sc.pp.neighbors(adata_, use_rep=emb_key)
multi_atlas_isolated_labels_f1_result["cell_subtype_zheng_2021"]["sample_name"][
emb_key
] = scib.me.isolated_labels_f1(
adata_,
label_key="cell_subtype_zheng_2021",
batch_key="sample_name",
embed=emb_key,
)
multi_atlas_isolated_labels_f1_result["cell_subtype_zheng_2021"]["study_name"][
emb_key
] = scib.me.isolated_labels_f1(
adata_,
label_key="cell_subtype_zheng_2021",
batch_key="study_name",
embed=emb_key,
)
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_chu_2023"].isin(["undefined"])
]
sc.pp.neighbors(adata_, use_rep=emb_key)
multi_atlas_isolated_labels_f1_result["cell_subtype_chu_2023"]["study_name"][
emb_key
] = scib.me.isolated_labels_f1(
adata_, label_key="cell_subtype_chu_2023", batch_key="study_name", embed=emb_key
)
Hyperparameter sensitivity analysis¶
Log or total variational normalization of the data¶
############################
# Log or total variational #
############################
for log_variational in [True, False]:
for total_variational in [True, False]:
vae_model = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_multi_atlas,
batch_key="sample_name",
additional_batch_keys=["study_name", "atlas_name"],
batch_embedding="embedding",
device="cuda:0",
label_key="cell_subtype_revision",
additional_label_keys=["cell_subtype_zheng_2021", "cell_subtype_chu_2023"],
log_variational=log_variational,
total_variational=total_variational,
)
vae_model.fit(max_epoch=10, lr=5e-5)
k = f"X_gex_log_variational_{log_variational}_total_variational_{total_variational}"
adata_cd8_multi_atlas.obsm[k] = vae_model.get_latent_embedding(
show_progress=True
)
np.save(
f"/slurm/home/yrd/liulab/xueziwei/2023-NM-Revision-GEX/huARdb_v2_GEX.CD8.hvg4k.pan_cancer_multi_atlas.{k}.npy",
adata_cd8_multi_atlas.obsm[k],
)
for log_variational in [True, False]:
for total_variational in [True, False]:
k = f"X_gex_log_variational_{log_variational}_total_variational_{total_variational}"
adata_cd8_multi_atlas.obsm[k] = np.load(
f"./huARdb_v2_GEX.CD8.hvg4k.pan_cancer_multi_atlas.{k}.npy"
)
hyper_parameter_variational_multi_atlas_graph_connectivity_result = {
"cell_subtype_revision": {},
"cell_subtype_zheng_2021": {},
"cell_subtype_chu_2023": {},
}
for log_variational in [True, False]:
for total_variational in [True, False]:
emb_key = f"X_gex_log_variational_{log_variational}_total_variational_{total_variational}"
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_revision"].isin(
["undefined", "???"]
)
]
sc.pp.neighbors(adata_, use_rep=emb_key)
hyper_parameter_variational_multi_atlas_graph_connectivity_result[
"cell_subtype_revision"
][emb_key] = scib.me.graph_connectivity(adata_, "cell_subtype_revision")
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_zheng_2021"].isin(["undefined"])
]
sc.pp.neighbors(adata_, use_rep=emb_key)
hyper_parameter_variational_multi_atlas_graph_connectivity_result[
"cell_subtype_zheng_2021"
][emb_key] = scib.me.graph_connectivity(
adata_,
label_key="cell_subtype_zheng_2021",
)
adata_ = adata_cd8_multi_atlas[
~adata_cd8_multi_atlas.obs["cell_subtype_chu_2023"].isin(["undefined"])
]
sc.pp.neighbors(adata_, use_rep=emb_key)
hyper_parameter_variational_multi_atlas_graph_connectivity_result[
"cell_subtype_chu_2023"
][emb_key] = scib.me.graph_connectivity(
adata_,
label_key="cell_subtype_chu_2023",
)
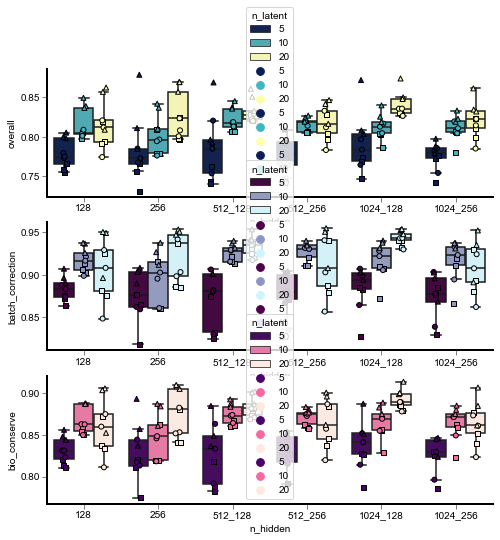
Hyperparameter set 2¶
for n_hidden in [[128], [256], [512, 128], [512, 256], [1024, 128], [1024, 256]]:
for n_latent in [5, 10, 20]:
for batch_hidden_dim in [16, 32, 64]:
vae_model = scatlasvae.model.scAtlasVAE(
adata=adata_cd8_multi_atlas,
hidden_stacks=n_hidden,
n_latent=n_latent,
batch_key="sample_name",
additional_batch_keys=["study_name", "atlas_name"],
batch_embedding="embedding",
batch_hidden_dim=batch_hidden_dim,
device="cuda:0",
label_key="cell_subtype_revision",
additional_label_keys=[
"cell_subtype_zheng_2021",
"cell_subtype_chu_2023",
],
)
vae_model.fit(max_epoch=10, lr=5e-5)
k = f'X_gex_n_hidden_{"_".join(list(map(str,n_hidden)))}_n_latent_{n_latent}_batch_hidden_dim_{batch_hidden_dim}'
adata_cd8_multi_atlas.obsm[k] = vae_model.get_latent_embedding(
show_progress=True
)
np.save(
f"./huARdb_v2_GEX.CD8.hvg4k.pan_cancer_multi_atlas.{k}.npy",
adata_cd8_multi_atlas.obsm[k],
)
Visualization of the Benchmark Result¶
from colour import Color
from matplotlib.colors import LinearSegmentedColormap
def make_colormap(colors, show_palette=False):
color_ramp = LinearSegmentedColormap.from_list(
"my_list", [Color(c1).rgb for c1 in colors]
)
if show_palette:
plt.figure(figsize=(15, 3))
plt.imshow(
[list(np.arange(0, len(colors), 0.1))],
interpolation="nearest",
origin="lower",
cmap=color_ramp,
)
plt.xticks([])
plt.yticks([])
return color_ramp
def reject_outliers(data, m=2):
return data[abs(data - np.mean(data)) < m * np.std(data)]
def rgb2hex(vals, rgbtype=1):
"""Converts RGB values in a variety of formats to Hex values.
@param vals An RGB/RGBA tuple
@param rgbtype Valid valus are:
1 - Inputs are in the range 0 to 1
256 - Inputs are in the range 0 to 255
@return A hex string in the form '#RRGGBB' or '#RRGGBBAA'"""
if len(vals) != 3 and len(vals) != 4:
raise Exception(
"RGB or RGBA inputs to RGBtoHex must have three or four elements!"
)
if rgbtype != 1 and rgbtype != 256:
raise Exception("rgbtype must be 1 or 256!")
# Convert from 0-1 RGB/RGBA to 0-255 RGB/RGBA
if rgbtype == 1:
vals = [255 * x for x in vals]
# Ensure values are rounded integers, convert to hex, and concatenate
return "#" + "".join(["{:02X}".format(int(round(x))) for x in vals])
metric_batch_colormap = make_colormap(
[
"#4D004A",
"#88419D",
"#8C96C6",
"#BFD3E6",
"#CEF5FD",
]
)
metric_bio_colormap = make_colormap(
[
"#49006A",
"#AE007E",
"#F768A1",
"#FCC5C0",
"#FFE9DE",
]
)
metrics_overall_colormap = make_colormap(
[
"#0A1D58",
"#235FA8",
"#40B6C4",
"#C7E9B4",
"#FDFFAB",
]
)
Extended Data Figure 3¶
import scanpy as sc
zheng_2021_annotation_cmap_cd8 = {
"CD8.c01.Tn.MAL": "#96C3D8",
"CD8.c02.Tm.IL7R": "#5D9BBE",
"CD8.c03.Tm.RPS12": "#F5B375",
"CD8.c04.Tm.CD52": "#C0937E",
"CD8.c05.Tem.CXCR5": "#67A59B",
"CD8.c06.Tem.GZMK": "#A4D38E",
"CD8.c07.Temra.CX3CR1": "#4A9D47",
"CD8.c08.Tk.TYROBP": "#F19294",
"CD8.c09.Tk.KIR2DL4": "#E45A5F",
"CD8.c10.Trm.ZNF683": "#3477A9",
"CD8.c11.Tex.PDCD1": "#BDA7CB",
"CD8.c12.Tex.CXCL13": "#684797",
"CD8.c13.Tex.myl12a": "#9983B7",
"CD8.c14.Tex.TCF7": "#CD9A99",
"CD8.c15.ISG.IFIT1": "#DD4B52",
"CD8.c16.MAIT.SLC4A10": "#DA8F6F",
"CD8.c17.Tm.NME1": "#F58135",
}
zheng_2021_annotation_cmap_cd4 = {
"CD4.c01.Tn.TCF7": "#78AECB",
"CD4.c02.Tn.PASK": "#639FB0",
"CD4.c03.Tn.ADSL": "#98C7A5",
"CD4.c04.Tn.il7r": "#83C180",
"CD4.c05.Tm.TNF": "#B2A4A5",
"CD4.c06.Tm.ANXA1": "#EC8D63",
"CD4.c07.Tm.ANXA2": "#CFC397",
"CD4.c08.Tm.CREM": "#F6B279",
"CD4.c09.Tm.CCL5": "#6197B4",
"CD4.c10.Tm.CAPG": "#CEA168",
"CD4.c11.Tm.GZMA": "#A0A783",
"CD4.c12.Tem.GZMK": "#9ACC90",
"CD4.c13.Temra.CX3CR1": "#6A9A52",
"CD4.c14.Th17.SLC4A10": "#E97679",
"CD4.c15.Th17.IL23R": "#DE4247",
"CD4.c16.Tfh.CXCR5": "#A38CBD",
"CD4.c17.TfhTh1.CXCL13": "#795FA3",
"CD4.c18.Treg.RTKN2": "#E0C880",
"CD4.c19.Treg.S1PR1": "#C28B65",
"CD4.c20.Treg.TNFRSF9": "#A65A34",
"CD4.c21.Treg.OAS1": "#DE4B3F",
"CD4.c22.ISG.IFIT1": "#DD9E82",
"CD4.c23.Mix.NME1": "#E78B75",
"CD4.c24.Mix.NME2": "#F7A96C",
"undefined": "#FFFFFF",
}
zheng_2021_annotation_cmap = zheng_2021_annotation_cmap_cd8.copy()
zheng_2021_annotation_cmap.update(zheng_2021_annotation_cmap_cd4)
chu_annotation_string = """
CD8-3 CD8_c3_Tn #E9ADC2
CD8-13 CD8_c13_Tn_TCF7 #AACC65
CD8-0 CD8_c0_Teff #00AFCA
CD8-2 CD8_c2_Teff #BBB7CB
CD8-8 CD8_c8_Teff_KLRG1 #E1A276
CD8-10 CD8_c10_Teff_CD244 #A5A2B3
CD8-11 CD8_c11_Teff_SEMA4A #A3AFA9
CD8-6 CD8_c6_Tcm #DD7A80
CD8-12 CD8_c12_Trm #A4BD83
CD8-7 CD8_c7_Tpex #EB9B7F
CD8-1 CD8_c1_Tex #76BCD8
CD8-4 CD8_c4_Tstr #E27C97
CD8-5 CD8_c5_Tisg #DF6C87
CD8-9 CD8_c9_Tsen #CCA891
CD4-2 CD4_c2_Tn #E0C8D9
CD4-6 CD4_c6_Tn_FHIT #F0A683
CD4-7 CD4_c7_Tn_TCEA3 #E5AE7C
CD4-9 CD4_c9_Tn_TCF7_SLC40A1 #A6AEBE
CD4-10 CD4_c10_Tn_LEF1_ANKRD55 #B3C28B
CD4-0 CD4_c0_Tcm #4CBBD2
CD4-5 CD4_c5_CTL #E9949E
CD4-1 CD4_c1_Treg #9FC6DB
CD4-3 CD4_c3_TFH #EFB3CC
CD4-8 CD4_c8_Th17 #C4ADA6
CD4-4 CD4_c4_Tstr #EF9AB9
CD4-11 CD4_c11_Tisg #C4D960
"""
subtype_color = {
"Tn": "#CEBF8F",
"Tcm": "#ffbb78",
"Early Tcm/Tem": "#ff7f0e",
"GZMK+ Tem": "#d62728",
"GNLY+ Temra": "#8c564b",
"CMC1+ Temra": "#e377c2",
"ZNF683+ Teff": "#6f3e7c",
"MAIT": "#17becf",
"ILTCK": "#aec7e8",
"ITGAE+ Trm": "#279e68",
"CREM+ Trm": "#aa40fc",
"ITGB2+ Trm": "#5ce041",
"Tpex": "#ff9896",
"GZMK+ Tex": "#C5B0D5",
"ITGAE+ Tex": "#C3823E",
"S100A11+ Tex": "#b5bd61",
"MACF1+ T": "#3288c9",
"Cycling T": "#f7b6d2",
}
subtype_color_alt = {"CD8+ " + k: v for k, v in subtype_color.items()}
chu_annotation = chu_annotation_string.split("\n")[1:-1]
chu_annotation = list(map(lambda x: x.split("\t"), chu_annotation))
import pandas as pd
chu_annotation = pd.DataFrame(chu_annotation)
chu_annotation_name = dict(zip(chu_annotation.iloc[:, 0], chu_annotation.iloc[:, 1]))
chu_annotation_cmap = dict(zip(chu_annotation.iloc[:, 1], chu_annotation.iloc[:, 2]))
chu_annotation_cmap_2 = dict(zip(chu_annotation.iloc[:, 0], chu_annotation.iloc[:, 2]))
default_20 = sc.pl.palettes.default_20
default_28 = sc.pl.palettes.default_28
godsnot_102 = sc.pl.palettes.godsnot_102
Extended Data Figure 3A¶
import os
import scanpy as sc
import warnings
warnings.filterwarnings("ignore")
adata_cd8_zheng_2021 = sc.read_h5ad("../data/adata_cd8_zheng_2021.h5ad")
for i in list(
filter(
lambda x: x.startswith("adata_cd8_zheng_2021.X") and x.endswith("umap.gpu.npy"),
os.listdir("embeddings"),
)
):
adata_cd8_zheng_2021.obsm[i.split("zheng_2021.")[1].split(".")[0]] = np.load(
os.path.join("embeddings", i)
)
zheng_2021_annotation_cmap_cd8["undefined"] = "#F7F7F7"
fig, axes = createSubplots(2, 6, figsize=(14, 4.5))
axes = axes.flatten()
for i, ax in zip(
[
"X_gex",
"X_gex_supervised",
"X_scVI",
"X_scPoli",
"X_scPoli_supervised",
"X_scANVI",
"X_SCALEX",
"X_scanorama",
"X_harmony",
"X_seuratv4_rpca",
"X_seuratv4_cca",
"X_pca",
],
axes,
):
if i in adata_cd8_zheng_2021.obsm.keys():
sc.pl.embedding(
adata_cd8_zheng_2021[
np.array(adata_cd8_zheng_2021.obsm[i][:, 0] < 24)
& np.array(adata_cd8_zheng_2021.obsm[i][:, 1] < 24)
],
color="cell_subtype_zheng_2021",
basis=i,
ax=ax,
show=False,
legend_loc="none",
palette=zheng_2021_annotation_cmap_cd8,
)
ax.set_title(i)
for ax in axes:
ax.set_xticks([])
ax.set_yticks([])
ax.spines["right"].set_color("none")
ax.spines["top"].set_color("none")
ax.spines["bottom"].set_color("none")
ax.spines["left"].set_color("none")
ax.set_title("")
ax.set_xlabel("")
ax.set_ylabel("")
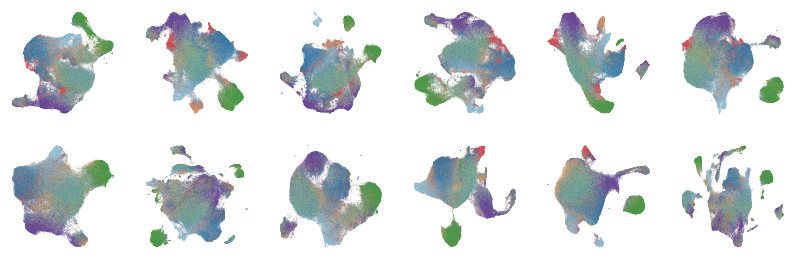

Extended Data Figure 3B¶
import warnings
warnings.filterwarnings("ignore")
fig, axes = createSubplots(2, 6, figsize=(14, 4.5))
axes = axes.flatten()
for i, ax in zip(
[
"X_gex",
"X_gex_supervised",
"X_scVI",
"X_scPoli",
"X_scPoli_supervised",
"X_scANVI",
"X_SCALEX",
"X_scanorama",
"X_harmony",
"X_seuratv4_rpca",
"X_seuratv4_cca",
"X_pca",
],
axes,
):
if i in adata_cd8_zheng_2021.obsm.keys():
sc.pl.embedding(
adata_cd8_zheng_2021[
np.array(adata_cd8_zheng_2021.obsm[i][:, 0] < 24)
& np.array(adata_cd8_zheng_2021.obsm[i][:, 1] < 24)
],
color="study_name",
basis=i,
ax=ax,
show=False,
legend_loc="none",
palette=default_28,
)
ax.set_title(i)
for ax in axes:
ax.set_xticks([])
ax.set_yticks([])
ax.spines["right"].set_color("none")
ax.spines["top"].set_color("none")
ax.spines["bottom"].set_color("none")
ax.spines["left"].set_color("none")
ax.set_title("")
ax.set_xlabel("")
ax.set_ylabel("")
WARNING: Length of palette colors is smaller than the number of categories (palette length: 28, categories length: 29. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 28, categories length: 29. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 28, categories length: 29. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 28, categories length: 29. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 28, categories length: 29. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 28, categories length: 29. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 28, categories length: 29. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 28, categories length: 29. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 28, categories length: 29. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 28, categories length: 29. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 28, categories length: 29. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 28, categories length: 29. Some categories will have the same color.
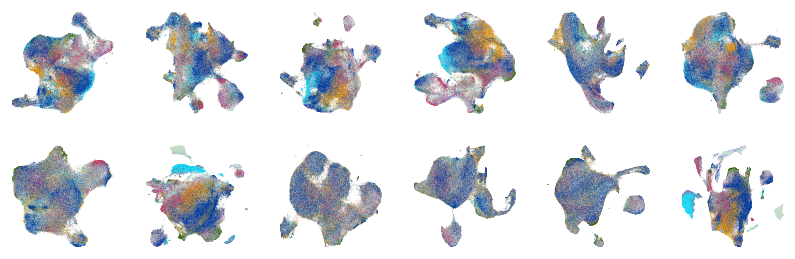

Extended Data Figure 3C¶
df = pd.read_csv("./Benchmark_zheng_2021.csv", index_col=0)
for i in range(df.shape[1]):
arr = df.iloc[:, i].to_numpy().flatten()
df.iloc[:, i] = (arr - arr.min()) / (arr.max() - arr.min())
metrics_batch = [
"silhouette_batch ASW (sample_name)",
"silhouette_batch ASW (study_name)",
"Graph connectivity",
"pcr_comparison (sample_name)",
"pcr_comparison (study_name)",
]
metrics_bio = [
"silhouette_score (label)",
"isolated_label_asw",
"isolated_label_f1",
]
df = df.loc[:, metrics_batch + metrics_bio]
df = df.iloc[::-1]
l = list(df.index)
gs_kw = dict(width_ratios=[1, 1, 1, 6])
fig, axes = plt.subplot_mosaic([[0, 1, 2, 3]], gridspec_kw=gs_kw, figsize=(10, 6))
mean_metrics_batch = df.loc[:, metrics_batch].mean(1)
rank = (1 - mean_metrics_batch).rank()
axes[1].barh(
y=list(range(df.shape[0])),
width=mean_metrics_batch,
color=list(
map(
lambda x: metric_batch_colormap(1 - x),
(rank - rank.min()) / (rank.max() - rank.min()),
)
),
)
mean_metrics_bio = df.loc[:, metrics_bio].mean(1)
rank = (1 - mean_metrics_bio).rank()
axes[2].barh(
y=list(range(df.shape[0])),
width=mean_metrics_bio,
color=list(
map(
lambda x: metric_bio_colormap(1 - x),
(rank - rank.min()) / (rank.max() - rank.min()),
)
),
)
mean_metrics_overall = df.mean(1)
rank = (1 - mean_metrics_overall).rank()
axes[0].barh(
y=list(range(df.shape[0])),
width=mean_metrics_overall,
color=list(
map(
lambda x: metrics_overall_colormap(1 - x),
(rank - rank.min()) / (rank.max() - rank.min()),
)
),
)
for i in range(df.shape[1]):
rank = (1 - df.iloc[:, i]).rank()
if df.columns[i] in metrics_batch:
c = list(
map(
lambda x: metric_batch_colormap(1 - x),
(rank - rank.min()) / (rank.max() - rank.min()),
)
)
else:
c = list(
map(
lambda x: metric_bio_colormap(1 - x),
(rank - rank.min()) / (rank.max() - rank.min()),
)
)
s = df.iloc[:, i]
s = reject_outliers(s)
s = (df.iloc[:, i] - s.min()) / (s.max() - s.min())
axes[3].scatter(
y=list(range(df.shape[0])),
x=[i] * df.shape[0],
s=np.array(s) * 400,
lw=0.1,
edgecolor="black",
c=c,
)
axes[3].set_yticks(range(df.shape[0]))
axes[3].set_yticklabels(df.index)
axes[3].set_xticks(range(df.shape[1]))
axes[3].set_xticklabels(df.columns, rotation=90)
axes[3].set_title("Extended Data Fig 3")
fig.savefig("./Benchmark_zheng_2021.pdf")
/opt/anaconda3/lib/python3.9/site-packages/matplotlib/collections.py:996: RuntimeWarning: invalid value encountered in sqrt
scale = np.sqrt(self._sizes) * dpi / 72.0 * self._factor
/opt/anaconda3/lib/python3.9/site-packages/matplotlib/collections.py:996: RuntimeWarning: invalid value encountered in sqrt
scale = np.sqrt(self._sizes) * dpi / 72.0 * self._factor

Extended Data Figure 4¶
Extended Data Figure 4A¶
import os
import scanpy as sc
import warnings
warnings.filterwarnings("ignore")
adata_cd8_chu = sc.read_h5ad("../data/adata_cd8_chu.h5ad")
for i in list(
filter(
lambda x: x.startswith("adata_cd8_chu.X") and x.endswith("umap.gpu.npy"),
os.listdir("embeddings"),
)
):
adata_cd8_chu.obsm[i.split("chu.")[1].split(".")[0]] = np.load(
os.path.join("embeddings", i)
)
zheng_2021_annotation_cmap_cd8["undefined"] = "#F7F7F7"
fig, axes = createSubplots(2, 6, figsize=(14, 4.5))
axes = axes.flatten()
for i, ax in zip(
[
"X_gex",
"X_gex_supervised",
"X_scVI",
"X_scPoli",
"X_scPoli_supervised",
"X_scANVI",
"X_SCALEX",
"X_scanorama",
"X_harmony",
"X_seuratv4_rpca",
"X_seuratv4_cca",
"X_pca",
],
axes,
):
if i in adata_cd8_chu.obsm.keys():
sc.pl.embedding(
adata_cd8_chu[
np.array(adata_cd8_chu.obsm[i][:, 0] < 24)
& np.array(adata_cd8_chu.obsm[i][:, 1] < 24)
],
color="cell_subtype_chu_2023",
basis=i,
ax=ax,
show=False,
legend_loc="none",
palette=chu_annotation_cmap_2,
)
for ax in axes:
ax.set_xticks([])
ax.set_yticks([])
ax.spines["right"].set_color("none")
ax.spines["top"].set_color("none")
ax.spines["bottom"].set_color("none")
ax.spines["left"].set_color("none")
ax.set_title("")
ax.set_xlabel("")
ax.set_ylabel("")
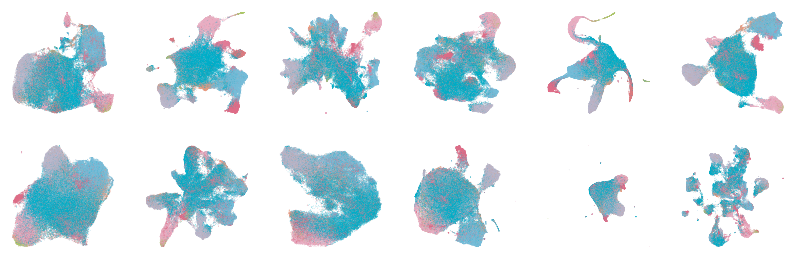

Extended Data Figure 4B¶
import os
import scanpy as sc
import warnings
warnings.filterwarnings("ignore")
adata_cd8_chu = sc.read_h5ad("../data/adata_cd8_chu.h5ad")
for i in list(
filter(
lambda x: x.startswith("adata_cd8_chu.X") and x.endswith("umap.gpu.npy"),
os.listdir("embeddings"),
)
):
adata_cd8_chu.obsm[i.split("chu.")[1].split(".")[0]] = np.load(
os.path.join("embeddings", i)
)
zheng_2021_annotation_cmap_cd8["undefined"] = "#F7F7F7"
fig, axes = createSubplots(2, 6, figsize=(14, 4.5))
axes = axes.flatten()
for i, ax in zip(
[
"X_gex",
"X_gex_supervised",
"X_scVI",
"X_scPoli",
"X_scPoli_supervised",
"X_scANVI",
"X_SCALEX",
"X_scanorama",
"X_harmony",
"X_seuratv4_rpca",
"X_seuratv4_cca",
"X_pca",
],
axes,
):
if i in adata_cd8_chu.obsm.keys():
sc.pl.embedding(
adata_cd8_chu[
np.array(adata_cd8_chu.obsm[i][:, 0] < 24)
& np.array(adata_cd8_chu.obsm[i][:, 1] < 24)
],
color="sample_name",
basis=i,
ax=ax,
show=False,
legend_loc="none",
palette=default_28,
)
for ax in axes:
ax.set_xticks([])
ax.set_yticks([])
ax.spines["right"].set_color("none")
ax.spines["top"].set_color("none")
ax.spines["bottom"].set_color("none")
ax.spines["left"].set_color("none")
ax.set_title("")
ax.set_xlabel("")
ax.set_ylabel("")
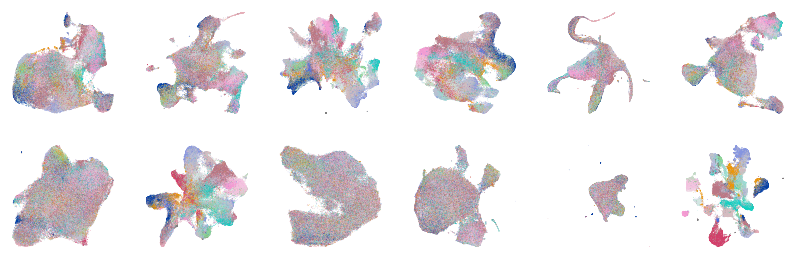

Extended Data Figure 4C¶
df = pd.read_csv("./benchmark_result_Chu_2023.csv", index_col=0)
for i in range(df.shape[1]):
arr = df.iloc[:, i].to_numpy().flatten()
df.iloc[:, i] = (arr - arr.min()) / (arr.max() - arr.min())
metrics_batch = [
"silhouette_batch ASW (study_name)",
"Graph connectivity",
"pcr_comparison",
]
metrics_bio = [
"silhouette_score (label)",
"isolated_label_asw (study_name)",
"isolated_label_f1 (study_name)",
]
df = df[df.index != "X_PCA"]
df = df.loc[:, metrics_batch + metrics_bio]
df = df.iloc[::-1]
l = list(df.index)
# l.remove("X_PCA")
# l = ['X_PCA'] + l
# df = df.loc[l]
gs_kw = dict(width_ratios=[1, 1, 1, 6])
fig, axes = plt.subplot_mosaic([[0, 1, 2, 3]], gridspec_kw=gs_kw, figsize=(10, 6))
mean_metrics_batch = df.loc[:, metrics_batch].mean(1)
rank = (1 - mean_metrics_batch).rank()
axes[1].barh(
y=list(range(df.shape[0])),
width=mean_metrics_batch,
color=list(
map(
lambda x: metric_batch_colormap(1 - x),
(rank - rank.min()) / (rank.max() - rank.min()),
)
),
)
mean_metrics_bio = df.loc[:, metrics_bio].mean(1)
rank = (1 - mean_metrics_bio).rank()
axes[2].barh(
y=list(range(df.shape[0])),
width=mean_metrics_bio,
color=list(
map(
lambda x: metric_bio_colormap(1 - x),
(rank - rank.min()) / (rank.max() - rank.min()),
)
),
)
mean_metrics_overall = df.mean(1)
rank = (1 - mean_metrics_overall).rank()
axes[0].barh(
y=list(range(df.shape[0])),
width=mean_metrics_overall,
color=list(
map(
lambda x: metrics_overall_colormap(1 - x),
(rank - rank.min()) / (rank.max() - rank.min()),
)
),
)
for i in range(df.shape[1]):
rank = (1 - df.iloc[:, i]).rank()
if df.columns[i] in metrics_batch:
c = list(
map(
lambda x: metric_batch_colormap(1 - x),
(rank - rank.min()) / (rank.max() - rank.min()),
)
)
else:
c = list(
map(
lambda x: metric_bio_colormap(1 - x),
(rank - rank.min()) / (rank.max() - rank.min()),
)
)
s = df.iloc[:, i]
s = reject_outliers(s)
s = (df.iloc[:, i] - s.min()) / (s.max() - s.min())
axes[3].scatter(
y=list(range(df.shape[0])),
x=[i] * df.shape[0],
s=np.array(s) * 400,
lw=0.1,
edgecolor="black",
c=c,
)
axes[3].set_yticks(range(df.shape[0]))
axes[3].set_yticklabels(df.index)
axes[3].set_xticks(range(df.shape[1]))
axes[3].set_xticklabels(df.columns, rotation=90)
axes[3].set_title("Extended Data Fig 4")
fig.savefig(".figures/benchmark_Chu_2023.pdf")

Extended Data Figure 5¶
for i in list(filter(lambda x: x.startswith("huARdb_v2_GEX.CD8.hvg4k.pan_cancer_multi_atlas.X") and x.endswith("umap.gpu.npy"), os.listdir("embeddings"))):
adata_cd8_multi_atlas.obsm[i.split("multi_atlas.")[1].split(".")[0]] = np.load(os.path.join("embeddings", i))
Extended Data Figure 5A¶
fig, axes = createSubplots(3, 6, figsize=(24, 14))
axes = axes.flatten()
for e, i in enumerate(
["X_gex", "X_scVI", "X_scPoli_supervised", "X_scANVI", "X_SCALEX", "X_pca"]
):
obsm = adata_cd8_multi_atlas.obsm[i]
for ax in axes[e * 3 : e * 3 + 3]:
ax.scatter(obsm[:, 0], obsm[:, 1], c="#E7E7E7", s=0.1, lw=0)
sc.pl.embedding(
adata_cd8_multi_atlas[
adata_cd8_multi_atlas.obs["atlas_name"].isin(["Chu_2023"])
],
color="cell_subtype_chu_2023",
ax=axes[e * 3 + 2],
palette=chu_annotation_cmap_2,
show=False,
basis=i,
legend_loc="none",
)
sc.pl.embedding(
adata_cd8_multi_atlas[
adata_cd8_multi_atlas.obs["atlas_name"].isin(["zheng_2021_PanCancer"])
],
color="cell_subtype_zheng_2021",
ax=axes[e * 3 + 1],
palette=zheng_2021_annotation_cmap_cd8,
show=False,
basis=i,
legend_loc="none",
)
sc.pl.embedding(
adata_cd8_multi_atlas[
np.array(adata_cd8_multi_atlas.obs["atlas_name"].isin(["huARdbv2"]))
],
color="cell_subtype_3",
ax=axes[e * 3 + 0],
palette=subtype_color,
show=False,
basis=i,
legend_loc="none",
)
for ax in axes:
ax.set_xticks([])
ax.set_yticks([])
ax.spines["right"].set_color("none")
ax.spines["top"].set_color("none")
ax.spines["bottom"].set_color("none")
ax.spines["left"].set_color("none")
ax.set_title("")
ax.set_xlabel("")
ax.set_ylabel("")
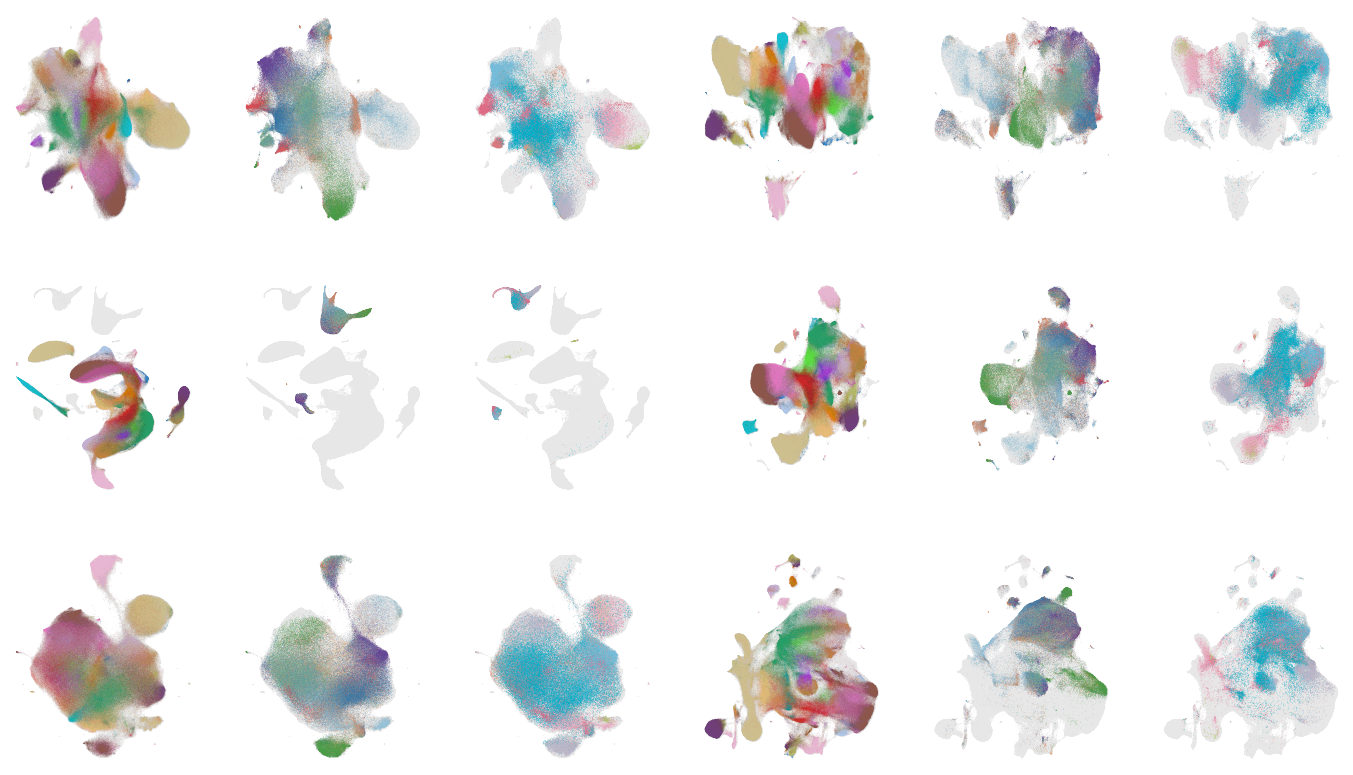

Extended Data Figure 5B¶
fig, axes = createSubplots(3, 6, figsize=(24, 14))
axes = axes.flatten()
for e, i in enumerate(
["X_gex", "X_scVI", "X_scPoli_supervised", "X_scANVI", "X_SCALEX", "X_pca"]
):
obsm = adata_cd8_multi_atlas.obsm[i]
for ax in axes[e * 3 : e * 3 + 3]:
ax.scatter(obsm[:, 0], obsm[:, 1], c="#E7E7E7", s=0.1, lw=0)
sc.pl.embedding(
adata_cd8_multi_atlas[
adata_cd8_multi_atlas.obs["atlas_name"].isin(["Chu_2023"])
],
color="study_name",
ax=axes[e * 3 + 2],
palette=default_20,
show=False,
basis=i,
legend_loc="none",
)
sc.pl.embedding(
adata_cd8_multi_atlas[
adata_cd8_multi_atlas.obs["atlas_name"].isin(["zheng_2021_PanCancer"])
],
color="study_name",
ax=axes[e * 3 + 1],
palette=default_28,
show=False,
basis=i,
legend_loc="none",
)
sc.pl.embedding(
adata_cd8_multi_atlas[
np.array(adata_cd8_multi_atlas.obs["atlas_name"].isin(["huARdbv2"]))
],
color="study_name",
ax=axes[e * 3 + 0],
palette=godsnot_102,
show=False,
basis=i,
legend_loc="none",
)
for ax in axes:
ax.set_xticks([])
ax.set_yticks([])
ax.spines["right"].set_color("none")
ax.spines["top"].set_color("none")
ax.spines["bottom"].set_color("none")
ax.spines["left"].set_color("none")
ax.set_title("")
ax.set_xlabel("")
ax.set_ylabel("")
WARNING: Length of palette colors is smaller than the number of categories (palette length: 20, categories length: 27. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 28, categories length: 29. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 20, categories length: 27. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 28, categories length: 29. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 20, categories length: 27. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 28, categories length: 29. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 20, categories length: 27. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 28, categories length: 29. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 20, categories length: 27. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 28, categories length: 29. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 20, categories length: 27. Some categories will have the same color.
WARNING: Length of palette colors is smaller than the number of categories (palette length: 28, categories length: 29. Some categories will have the same color.
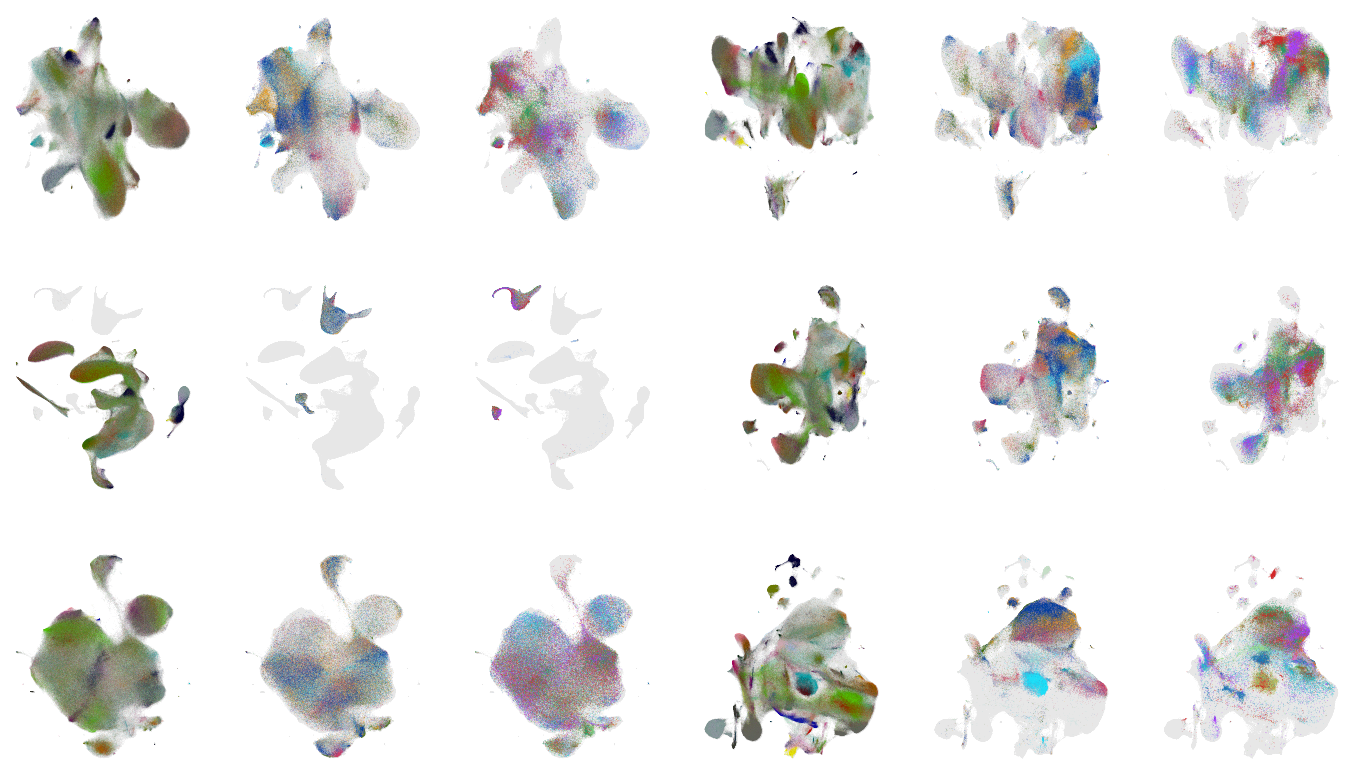

Extended Data Figure 6¶
Extended Data Figure 6D¶
df_all = pd.read_csv("./Benchmark_hyperparameter_all.csv")
df_all["n_hidden_int"] = df_all["n_hidden"].apply(
lambda x: (int(x.split("_")[0]), int(x.split("_")[-1]))
)
metrics_batch_colormap = [
"#4D004A",
"#8C96C6",
"#CEF5FD",
]
metrics_bio_colormap = [
"#49006A",
"#F768A1",
"#FFE9DE",
]
metrics_overall_colormap = [
"#0A1D58",
"#40B6C4",
"#FDFFAB",
]
fig, ax = createSubplots(3, 1)
markdict = {
"cell_subtype_revision": "s",
"cell_subtype_zheng_2021": "o",
"cell_subtype_chu_2023": "^",
}
sns.boxplot(
data=df_all.sort_values("n_hidden_int"),
x="n_hidden",
y="overall",
hue="n_latent",
ax=ax[0],
showfliers=False,
palette=metrics_overall_colormap,
)
for k, v in markdict.items():
sns.stripplot(
data=df_all[df_all["atlas_name"] == k].sort_values("n_hidden_int"),
x="n_hidden",
y="overall",
hue="n_latent",
ax=ax[0],
dodge=True,
marker=v,
edgecolor="black",
linewidth=1,
palette=metrics_overall_colormap,
)
sns.boxplot(
data=df_all.sort_values("n_hidden_int"),
x="n_hidden",
y="batch_correction",
hue="n_latent",
ax=ax[1],
showfliers=False,
palette=metrics_batch_colormap,
)
for k, v in markdict.items():
sns.stripplot(
data=df_all[df_all["atlas_name"] == k].sort_values("n_hidden_int"),
x="n_hidden",
y="batch_correction",
hue="n_latent",
ax=ax[1],
dodge=True,
marker=v,
edgecolor="black",
linewidth=1,
palette=metrics_batch_colormap,
)
sns.boxplot(
data=df_all.sort_values("n_hidden_int"),
x="n_hidden",
y="bio_conserve",
hue="n_latent",
ax=ax[2],
showfliers=False,
palette=metrics_bio_colormap,
)
for k, v in markdict.items():
sns.stripplot(
data=df_all[df_all["atlas_name"] == k].sort_values("n_hidden_int"),
x="n_hidden",
y="bio_conserve",
hue="n_latent",
ax=ax[2],
dodge=True,
marker=v,
edgecolor="black",
linewidth=1,
palette=metrics_bio_colormap,
)